Mutant p53 is a transcriptional co-factor that binds to G-rich regulatory regions of active genes and generates transcriptional plasticity
- PMID: 22894900
- PMCID: PMC3466528
- DOI: 10.4161/cc.21646
Mutant p53 is a transcriptional co-factor that binds to G-rich regulatory regions of active genes and generates transcriptional plasticity
Abstract
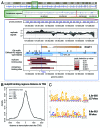
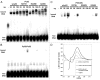
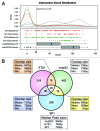
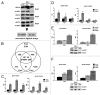
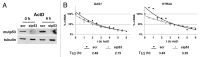
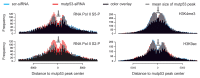
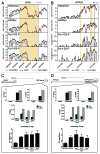
The molecular mechanisms underlying mutant p53 (mutp53) "gain-of-function" (GOF) are still insufficiently understood, but there is evidence that mutp53 is a transcriptional regulator that is recruited by specialized transcription factors. Here we analyzed the binding sites of mutp53 and the epigenetic status of mutp53-regulated genes that had been identified by global expression profiling upon depletion of endogenous mutp53 (R273H) expression in U251 glioblastoma cells. We found that mutp53 preferentially and autonomously binds to G/C-rich DNA around transcription start sites (TSS) of many genes characterized by active chromatin marks (H3K4me3) and frequently associated with transcription-competent RNA polymerase II. Mutp53-bound regions overlap predominantly with CpG islands and are enriched in G4-motifs that are prone to form G-quadruplex structures. In line, mutp53 binds and stabilizes a well-characterized G-quadruplex structure in vitro. Hence, we assume that binding of mutp53 to G/C-rich DNA regions associated with a large set of cancer-relevant genes is an initial step in their regulation by mutp53. Using GAS1 and HTR2A as model genes, we show that mutp53 affects several parameters of active transcription. Finally, we discuss a dual mode model of mutp53 GOF, which includes both stochastic and deterministic components.
Figures
References
-
- Brosh R, Rotter V. When mutants gain new powers: news from the mutant p53 field. Nat Rev Cancer. 2009;9:701–13. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous
