Enterohemorrhagic and enteropathogenic Escherichia coli evolved different strategies to resist antimicrobial peptides
- PMID: 22895086
- PMCID: PMC3495793
- DOI: 10.4161/gmic.21656
Enterohemorrhagic and enteropathogenic Escherichia coli evolved different strategies to resist antimicrobial peptides
Abstract
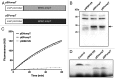
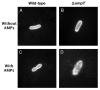
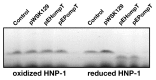
Enterohemorrhagic and enteropathogenic Escherichia coli (EHEC and EPEC) are enteric human pathogens that colonize the large and small intestines, respectively. To establish infection EHEC and EPEC must overcome innate host defenses, such as antimicrobial peptides (AMPs) produced by the intestinal epithelium. Gram-negative pathogens have evolved different mechanisms to resist AMPs, including outer-membrane proteases that degrade AMPs. We showed that the protease OmpT degrades the human AMP LL-37 more rapidly in EHEC than in EPEC. Promoter-swap experiments showed that this is due to differences in the promoters of the two genes, leading to greater ompT expression and subsequently greater levels of OmpT in EHEC. Here, we propose that the different ompT expression in EHEC and EPEC reflects the varying levels of LL-37 throughout the human intestinal tract. These data suggest that EHEC and EPEC adapted to their specific niches by developing distinct AMP-specific resistance mechanisms.
Figures
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
