Recent advances in mapping environmental microbial metabolisms through 13C isotopic fingerprints
- PMID: 22896564
- PMCID: PMC3479925
- DOI: 10.1098/rsif.2012.0396
Recent advances in mapping environmental microbial metabolisms through 13C isotopic fingerprints
Abstract
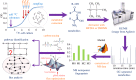
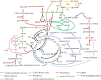
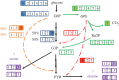
After feeding microbes with a defined (13)C substrate, unique isotopic patterns (isotopic fingerprints) can be formed in their metabolic products. Such labelling information not only can provide novel insights into functional pathways but also can determine absolute carbon fluxes through the metabolic network via metabolic modelling approaches. This technique has been used for finding pathways that may have been mis-annotated in the past, elucidating new enzyme functions, and investigating cell metabolisms in microbial communities. In this review paper, we summarize the applications of (13)C approaches to analyse novel cell metabolisms for the past 3 years. The isotopic fingerprints (defined as unique isotopomers useful for pathway identifications) have revealed the operations of the Entner-Doudoroff pathway, the reverse tricarboxylic acid cycle, new enzymes for biosynthesis of central metabolites, diverse respiration routes in phototrophic metabolism, co-metabolism of carbon nutrients and novel CO(2) fixation pathways. This review also discusses new isotopic methods to map carbon fluxes in global metabolisms, as well as potential factors influencing the metabolic flux quantification (e.g. metabolite channelling, the isotopic purity of (13)C substrates and the isotopic effect). Although (13)C labelling is not applicable to all biological systems (e.g. microbial communities), recent studies have shown that this method has a significant value in functional characterization of poorly understood micro-organisms, including species relevant for biotechnology and human health.
Figures
References
-
- Zamboni N., Fendt S. M., Rühl M., Sauer U. 2009. 13C-based metabolic flux analysis. Nat. Protoc. 4, 878–892 10.1038/nprot.2009.58 (doi:10.1038/nprot.2009.58) - DOI - PubMed
-
- Zamboni N., Sauer U. 2009. Novel biological insights through metabolomics and 13C-flux analysis. Curr. Opin. Microbiol. 12, 553–558 10.1016/j.mib.2009.08.003 (doi:10.1016/j.mib.2009.08.003) - DOI - PubMed
-
- Wiechert W. 2001. 13C metabolic flux analysis. Metab. Eng. 3, 195–206 10.1006/mben.2001.0187 (doi:10.1006/mben.2001.0187) - DOI - PubMed
-
- Wittmann C. 2007. Fluxome analysis using GC-MS. Microb. Cell Factories 6, 6. 10.1186/1475-2859-6-6 (doi:10.1186/1475-2859-6-6) - DOI - PMC - PubMed
-
- Wittmann C. 2002. Metabolic flux analysis using mass spectrometry. Adv. Biochem. Eng./Biotechnol. 74, 39–64 10.1007/3-540-45736-4_3 (doi:10.1007/3-540-45736-4_3) - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous
