Evaluation of phosphatidylinositol-4-kinase IIIα as a hepatitis C virus drug target
- PMID: 22896614
- PMCID: PMC3486294
- DOI: 10.1128/JVI.01320-12
Evaluation of phosphatidylinositol-4-kinase IIIα as a hepatitis C virus drug target
Abstract
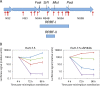
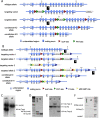
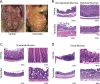
Phosphatidylinositol-4-kinase IIIα (PI4KIIIα) is an essential host cell factor for hepatitis C virus (HCV) replication. An N-terminally truncated 130-kDa form was used to reconstitute an in vitro biochemical lipid kinase assay that was optimized for small-molecule compound screening and identified potent and specific inhibitors. Cell culture studies with PI4KIIIα inhibitors demonstrated that the kinase activity was essential for HCV RNA replication. Two PI4KIIIα inhibitors were used to select cell lines harboring HCV replicon mutants with a 20-fold loss in sensitivity to the compounds. Reverse genetic mapping isolated an NS4B-NS5A segment that rescued HCV RNA replication in PIK4IIIα-deficient cells. HCV RNA replication occurs on specialized membranous webs, and this study with PIK4IIIα inhibitor-resistant mutants provides a genetic link between NS4B/NS5A functions and PI4-phosphate lipid metabolism. A comprehensive assessment of PI4KIIIα as a drug target included its evaluation for pharmacologic intervention in vivo through conditional transgenic murine lines that mimic target-specific inhibition in adult mice. Homozygotes that induce a knockout of the kinase domain or knock in a single amino acid substitution, kinase-defective PI4KIIIα, displayed a lethal phenotype with a fairly widespread mucosal epithelial degeneration of the gastrointestinal tract. This essential host physiologic role raises doubt about the pursuit of PI4KIIIα inhibitors for treatment of chronic HCV infection.
Figures




Similar articles
-
Hepatitis C Virus Subverts Human Choline Kinase-α To Bridge Phosphatidylinositol-4-Kinase IIIα (PI4KIIIα) and NS5A and Upregulates PI4KIIIα Activation, Thereby Promoting the Translocation of the Ternary Complex to the Endoplasmic Reticulum for Viral Replication.J Virol. 2017 Jul 27;91(16):e00355-17. doi: 10.1128/JVI.00355-17. Print 2017 Aug 15. J Virol. 2017. PMID: 28566381 Free PMC article.
-
Metabolism of phosphatidylinositol 4-kinase IIIα-dependent PI4P Is subverted by HCV and is targeted by a 4-anilino quinazoline with antiviral activity.PLoS Pathog. 2012;8(3):e1002576. doi: 10.1371/journal.ppat.1002576. Epub 2012 Mar 8. PLoS Pathog. 2012. PMID: 22412376 Free PMC article.
-
Mapping of functional domains of the lipid kinase phosphatidylinositol 4-kinase type III alpha involved in enzymatic activity and hepatitis C virus replication.J Virol. 2014 Sep 1;88(17):9909-26. doi: 10.1128/JVI.01063-14. Epub 2014 Jun 11. J Virol. 2014. PMID: 24920820 Free PMC article.
-
Antiviral activity and resistance of HCV NS5A replication complex inhibitors.Curr Opin Virol. 2013 Oct;3(5):514-20. doi: 10.1016/j.coviro.2013.06.014. Epub 2013 Jul 27. Curr Opin Virol. 2013. PMID: 23896281 Review.
-
Host cell kinases and the hepatitis C virus life cycle.Biochim Biophys Acta. 2015 Oct;1854(10 Pt B):1657-62. doi: 10.1016/j.bbapap.2015.04.011. Epub 2015 Apr 18. Biochim Biophys Acta. 2015. PMID: 25896387 Review.
Cited by
-
A Forward Genetic Screen Targeting the Endothelium Reveals a Regulatory Role for the Lipid Kinase Pi4ka in Myelo- and Erythropoiesis.Cell Rep. 2018 Jan 30;22(5):1211-1224. doi: 10.1016/j.celrep.2018.01.017. Cell Rep. 2018. PMID: 29386109 Free PMC article.
-
Host-targeting agents in the treatment of hepatitis C: a beginning and an end?Antiviral Res. 2013 Nov;100(2):555-61. doi: 10.1016/j.antiviral.2013.09.020. Epub 2013 Sep 30. Antiviral Res. 2013. PMID: 24091203 Free PMC article. Review.
-
Viral rewiring of cellular lipid metabolism to create membranous replication compartments.Curr Opin Cell Biol. 2017 Aug;47:24-33. doi: 10.1016/j.ceb.2017.02.005. Epub 2017 Feb 24. Curr Opin Cell Biol. 2017. PMID: 28242560 Free PMC article. Review.
-
Biallelic PI4KA variants cause neurological, intestinal and immunological disease.Brain. 2021 Dec 31;144(12):3597-3610. doi: 10.1093/brain/awab313. Brain. 2021. PMID: 34415310 Free PMC article.
-
Two Novel Variants in PI4KA in a Family Presenting With Hereditary Spastic Paraparesis: A Case Report.Neurol Genet. 2024 Apr 25;10(3):e200152. doi: 10.1212/NXG.0000000000200152. eCollection 2024 Jun. Neurol Genet. 2024. PMID: 38685974 Free PMC article.
References
-
- Ahn J, et al. 2004. Systematic identification of hepatocellular proteins interacting with NS5A of the hepatitis C virus. J. Biochem. Mol. Biol. 37:741–748 - PubMed
-
- Akimana C, Al-Khodor S, Abu Kwaik Y. 2010. Host factors required for modulation of phagosome biogenesis and proliferation of Francisella tularensis within the cytosol. PLoS One 5:e11025 doi:10.1371/journal.pone.0011025 - DOI - PMC - PubMed
-
- Balla A, Balla T. 2006. Phosphatidylinositol 4-kinases: old enzymes with emerging functions. Trends Cell Biol. 16:351–361 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

