Cohesin acetylation promotes sister chromatid cohesion only in association with the replication machinery
- PMID: 22896698
- PMCID: PMC3464539
- DOI: 10.1074/jbc.M112.400192
Cohesin acetylation promotes sister chromatid cohesion only in association with the replication machinery
Abstract
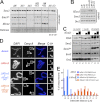
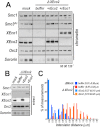
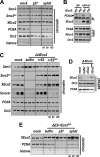
Acetylation of the Smc3 subunit of cohesin is essential to establish functional cohesion between sister chromatids. Smc3 acetylation is catalyzed by members of the Eco family of acetyltransferases, although the mechanism by which acetylation is regulated and how it promotes cohesion are largely unknown. In vertebrates, the cohesin complex binds to chromatin during mitotic exit and is converted to a functional form during or shortly after DNA replication. The conserved proliferating cell nuclear antigen-interacting protein box motif in yeast Eco1 is required for function, and cohesin is acetylated during the S phase. This has led to the notion that acetylation of cohesin is stimulated by interaction of Eco1 with the replication machinery. Here we show that in vertebrates Smc3 acetylation occurs independently of DNA replication. Smc3 is readily acetylated before replication is initiated and after DNA replication is complete. However, we also show that functional acetylation occurs only in association with the replication machinery: disruption of the interaction between XEco2 and proliferating cell nuclear antigen prevents cohesion establishment while having little impact on the overall levels of Smc3 acetylation. These results demonstrate that Smc3 acetylation can occur throughout interphase but that only acetylation in association with the replication fork promotes sister chromatid cohesion. These data reveal how the generation of cohesion is limited to the appropriate time and place during the cell cycle and provide insight into the mechanism by which acetylation ensures cohesion.
Figures




References
-
- Rolef Ben-Shahar T., Heeger S., Lehane C., East P., Flynn H., Skehel M., Uhlmann F. (2008) Eco1-dependent cohesin acetylation during establishment of sister chromatid cohesion. Science 321, 563–566 - PubMed
-
- Rowland B. D., Roig M. B., Nishino T., Kurze A., Uluocak P., Mishra A., Beckouët F., Underwood P., Metson J., Imre R., Mechtler K., Katis V. L., Nasmyth K. (2009) Building sister chromatid cohesion. Smc3 acetylation counteracts an antiestablishment activity. Mol. Cell 33, 763–774 - PubMed
-
- Zhang J., Shi X., Li Y., Kim B. J., Jia J., Huang Z., Yang T., Fu X., Jung S. Y., Wang Y., Zhang P., Kim S. T., Pan X., Qin J. (2008) Acetylation of Smc3 by Eco1 is required for S phase sister chromatid cohesion in both human and yeast. Mol. Cell 31, 143–151 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

