Radical SAM enzymes involved in the biosynthesis of purine-based natural products
- PMID: 22902275
- PMCID: PMC4022190
- DOI: 10.1016/j.bbapap.2012.07.014
Radical SAM enzymes involved in the biosynthesis of purine-based natural products
Abstract
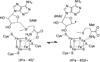
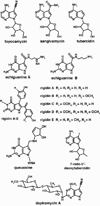
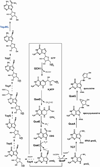
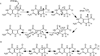
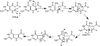
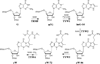
The radical S-adenosyl-l-methionine (SAM) superfamily is a widely distributed group of iron-sulfur containing proteins that exploit the reactivity of the high energy intermediate, 5'-deoxyadenosyl radical, which is produced by the reductive cleavage of SAM, to carry-out complex radical-mediated transformations. The reactions catalyzed by radical SAM enzymes range from simple group migrations to complex reactions in protein and RNA modification. This review will highlight three radical SAM enzymes that catalyze reactions involving modified guanosines in the biosynthesis pathways of the hypermodified tRNA base wybutosine; secondary metabolites of 7-deazapurine structure, including the hypermodified tRNA base queuosine; and the redox cofactor F(420). This article is part of a Special Issue entitled: Radical SAM enzymes and Radical Enzymology.
Copyright © 2012 Elsevier B.V. All rights reserved.
Figures
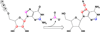
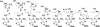
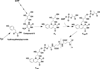
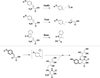
References
-
- Sofia H, Chen G, Hetzler B, Reyes-Spindola J, Miller N. Radical SAM, a novel protein superfamily linking unresolved steps in familiar biosynthetic pathways with radical mechanisms: functional characterization using new analysis and information visualization methods. Nucleic Acids Res. 2001;29:1097–1106. - PMC - PubMed
-
- Walsby CJ, Ortillo D, Broderick WE, Broderick JB, Hoffman BM. An Anchoring Role for FeS Clusters: Chelation of the Amino Acid Moiety of S-Adenosylmethionine to the Unique Iron Site of the [4Fe-4S]. Cluster of Pyruvate Formate-Lyase Activating Enzyme. J Am Chem Soc. 2002;124:11270–11271. - PubMed
-
- Walsby CJ, Ortillo D, Yang J, Nnyepi MR, Broderick WE, Hoffman BM, et al. Spectroscopic Approaches to Elucidating Novel Iron-Sulfur Chemistry in the ”Radical-SAM” Protein Superfamily. Inorg. Chem. 2005;44:727–741. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

