Deletion of the Aspergillus flavus orthologue of A. nidulans fluG reduces conidiation and promotes production of sclerotia but does not abolish aflatoxin biosynthesis
- PMID: 22904054
- PMCID: PMC3485703
- DOI: 10.1128/AEM.01241-12
Deletion of the Aspergillus flavus orthologue of A. nidulans fluG reduces conidiation and promotes production of sclerotia but does not abolish aflatoxin biosynthesis
Abstract
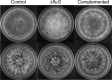
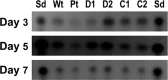
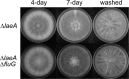
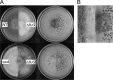
The fluG gene is a member of a family of genes required for conidiation and sterigmatocystin production in Aspergillus nidulans. We examined the role of the Aspergillus flavus fluG orthologue in asexual development and aflatoxin biosynthesis. Deletion of fluG in A. flavus yielded strains with an approximately 3-fold reduction in conidiation but a 30-fold increase in sclerotial formation when grown on potato dextrose agar in the dark. The concurrent developmental changes suggest that A. flavus FluG exerts opposite effects on a mutual signaling pathway for both processes. The altered conidial development was in part attributable to delayed expression of brlA, a gene controlling conidiophore formation. Unlike the loss of sterigmatocystin production by A. nidulans fluG deletion strains, aflatoxin biosynthesis was not affected by the fluG deletion in A. flavus. In A. nidulans, FluG was recently found to be involved in the formation of dehydroaustinol, a component of a diffusible signal of conidiation. Coculturing experiments did not show a similar diffusible meroterpenoid secondary metabolite produced by A. flavus. These results suggest that the function of fluG and the signaling pathways related to conidiation are different in the two related aspergilli.
Figures

References
-
- Adams TH, Boylan MT, Timberlake WE. 1988. brlA is necessary and sufficient to direct conidiophore development in Aspergillus nidulans. Cell 54:353–362 - PubMed
-
- Aguirre J. 1993. Spatial and temporal controls of the Aspergillus brlA developmental regulatory gene. Mol. Microbiol. 8:211–218 - PubMed
-
- Bayram O, et al. 2008. VelB/VeA/LaeA complex coordinates light signal with fungal development and secondary metabolism. Science 320:1504–1506 - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

