Oxa1-ribosome complexes coordinate the assembly of cytochrome C oxidase in mitochondria
- PMID: 22904327
- PMCID: PMC3464553
- DOI: 10.1074/jbc.M112.382630
Oxa1-ribosome complexes coordinate the assembly of cytochrome C oxidase in mitochondria
Abstract
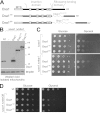
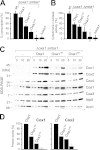
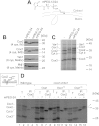
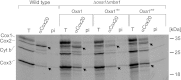
The terminal enzyme of the respiratory chain, cytochrome c oxidase, consists of a hydrophobic reaction center formed by three mitochondrially encoded subunits with which 9-10 nuclear encoded subunits are associated. The three core subunits are synthesized on mitochondrial ribosomes and inserted into the inner membrane in a co-translational reaction facilitated by the Oxa1 insertase. Oxa1 consists of an N-terminal insertase domain and a C-terminal ribosome-binding region. Mutants lacking the C-terminal region show specific defects in co-translational insertion, suggesting that the close contact of the ribosome with the insertase promotes co-translational insertion of nascent chains. In this study, we inserted flexible linkers of 100 or 200 amino acid residues between the insertase domain and ribosome-binding region of Oxa1 of Saccharomyces cerevisiae. In the absence of the ribosome receptor Mba1, these linkers caused a length-dependent decrease in mitochondrial respiratory activity caused by diminished levels of cytochrome c oxidase. Interestingly, considerable amounts of mitochondrial translation products were still integrated into the inner membrane in these linker mutants. However, they showed severe defects in later stages of the biogenesis process, presumably during assembly into functional complexes. Our observations suggest that the close proximity of Oxa1 to ribosomes is not only used to improve membrane insertion but is also critical for the productive assembly of the subunits of the cytochrome c oxidase. This points to a role for Oxa1 in the spatial coordination of the ribosome with assembly factors that are critical for enzyme biogenesis.
Figures

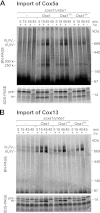

References
-
- Young J. C., Agashe V. R., Siegers K., Hartl F. U. (2004) Pathways of chaperone-mediated protein folding in the cytosol. Nat. Rev. Mol. Cell Biol. 5, 781–791 - PubMed
-
- Kramer G., Boehringer D., Ban N., Bukau B. (2009) The ribosome as a platform for co-translational processing, folding, and targeting of newly synthesized proteins. Nat. Struct. Mol. Biol. 16, 589–597 - PubMed
-
- Giglione C., Fieulaine S., Meinnel T. (2009) Co-translational processing mechanisms: towards a dynamic 3D model. Trends Biochem. Sci. 34, 417–426 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

