Terpenoid synthase structures: a so far incomplete view of complex catalysis
- PMID: 22907771
- PMCID: PMC3448952
- DOI: 10.1039/c2np20059g
Terpenoid synthase structures: a so far incomplete view of complex catalysis
Abstract
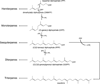
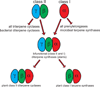
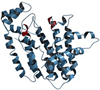
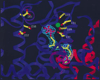
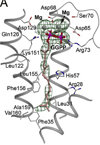
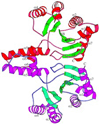
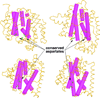
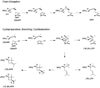
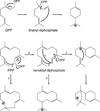
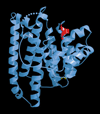
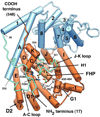
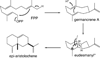
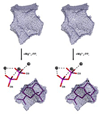
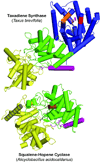
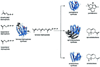
The complexity of terpenoid natural products has drawn significant interest, particularly since their common (poly)isoprenyl origins were discovered. Notably, much of this complexity is derived from the highly variable cyclized and/or rearranged nature of the observed hydrocarbon skeletal structures. Indeed, at least in some cases it is difficult to immediately recognize their derivation from poly-isoprenyl precursors. Nevertheless, these diverse structures are formed by sequential elongation to acyclic precursors, most often with subsequent cyclization and/or rearrangement. Strikingly, the reactions used to assemble and diversify terpenoid backbones share a common carbocationic driven mechanism, although the means by which the initial carbocation is generated does vary. High-resolution crystal structures have been obtained for at least representative examples from each of the various types of enzymes involved in producing terpenoid hydrocarbon backbones. However, while this has certainly led to some insights into the enzymatic structure-function relationships underlying the elongation and simpler cyclization reactions, our understanding of the more complex cyclization and/or rearrangement reactions remains limited. Accordingly, selected examples are discussed here to demonstrate our current understanding, its limits, and potential ways forward.
Figures
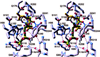



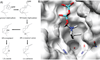
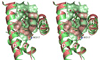



Similar articles
-
Structural and Chemical Biology of Terpenoid Cyclases.Chem Rev. 2017 Sep 13;117(17):11570-11648. doi: 10.1021/acs.chemrev.7b00287. Epub 2017 Aug 25. Chem Rev. 2017. PMID: 28841019 Free PMC article. Review.
-
Unearthing the roots of the terpenome.Curr Opin Chem Biol. 2008 Apr;12(2):141-50. doi: 10.1016/j.cbpa.2007.12.008. Epub 2008 Feb 20. Curr Opin Chem Biol. 2008. PMID: 18249199 Free PMC article. Review.
-
Terpenoid biosynthesis off the beaten track: unconventional cyclases and their impact on biomimetic synthesis.Angew Chem Int Ed Engl. 2015 Feb 23;54(9):2604-26. doi: 10.1002/anie.201407883. Epub 2014 Dec 8. Angew Chem Int Ed Engl. 2015. PMID: 25488271 Review.
-
Assembly-Line Catalysis in Bifunctional Terpene Synthases.Acc Chem Res. 2021 Oct 19;54(20):3780-3791. doi: 10.1021/acs.accounts.1c00296. Epub 2021 Jul 13. Acc Chem Res. 2021. PMID: 34254507 Free PMC article. Review.
-
Managing and manipulating carbocations in biology: terpenoid cyclase structure and mechanism.Curr Opin Struct Biol. 1998 Dec;8(6):695-703. doi: 10.1016/s0959-440x(98)80088-2. Curr Opin Struct Biol. 1998. PMID: 9914250 Review.
Cited by
-
The biosynthetic origin of irregular monoterpenes in Lavandula: isolation and biochemical characterization of a novel cis-prenyl diphosphate synthase gene, lavandulyl diphosphate synthase.J Biol Chem. 2013 Mar 1;288(9):6333-41. doi: 10.1074/jbc.M112.431171. Epub 2013 Jan 10. J Biol Chem. 2013. PMID: 23306202 Free PMC article.
-
The application of synthetic biology to elucidation of plant mono-, sesqui-, and diterpenoid metabolism.Mol Plant. 2015 Jan;8(1):6-16. doi: 10.1016/j.molp.2014.12.002. Epub 2014 Dec 11. Mol Plant. 2015. PMID: 25578268 Free PMC article. Review.
-
Identification and Characterization of Bifunctional Drimenol Synthases of Marine Bacterial Origin.ACS Chem Biol. 2022 May 20;17(5):1226-1238. doi: 10.1021/acschembio.2c00163. Epub 2022 Apr 21. ACS Chem Biol. 2022. PMID: 35446557 Free PMC article.
-
In planta and in silico characterization of five sesquiterpene synthases from Vitis vinifera (cv. Shiraz) berries.Planta. 2019 Jan;249(1):59-70. doi: 10.1007/s00425-018-2986-7. Epub 2018 Aug 22. Planta. 2019. PMID: 30136197
-
Crystal structure of F95Q epi-isozizaene synthase, an engineered sesquiterpene cyclase that generates biofuel precursors β- and γ-curcumene.J Struct Biol. 2019 Aug 1;207(2):218-224. doi: 10.1016/j.jsb.2019.05.011. Epub 2019 May 29. J Struct Biol. 2019. PMID: 31152775 Free PMC article.
References
-
- Croteau R, Kutchan TM, Lewis NG. In: Biochemistry & Molecular Biology of Plants. Buchanan B, Gruissem W, Jones R, editors. Rockville, MD, USA: Am. Soc. Plant Biologists; 2000. pp. 1250–1318.
-
- Wendt KU, Schulz GE. Structure. 1998;6:127–133. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

