Broad-spectrum antivirals against 3C or 3C-like proteases of picornaviruses, noroviruses, and coronaviruses
- PMID: 22915796
- PMCID: PMC3486288
- DOI: 10.1128/JVI.01348-12
Broad-spectrum antivirals against 3C or 3C-like proteases of picornaviruses, noroviruses, and coronaviruses
Abstract
Phylogenetic analysis has demonstrated that some positive-sense RNA viruses can be classified into the picornavirus-like supercluster, which includes picornaviruses, caliciviruses, and coronaviruses. These viruses possess 3C or 3C-like proteases (3Cpro or 3CLpro, respectively), which contain a typical chymotrypsin-like fold and a catalytic triad (or dyad) with a Cys residue as a nucleophile. The conserved key sites of 3Cpro or 3CLpro may serve as attractive targets for the design of broad-spectrum antivirals for multiple viruses in the supercluster. We previously reported the structure-based design and synthesis of potent protease inhibitors of Norwalk virus (NV), a member of the Caliciviridae family. We report herein the broad-spectrum antiviral activities of three compounds possessing a common dipeptidyl residue with different warheads, i.e., an aldehyde (GC373), a bisulfite adduct (GC376), and an α-ketoamide (GC375), against viruses that belong to the supercluster. All compounds were highly effective against the majority of tested viruses, with half-maximal inhibitory concentrations in the high nanomolar or low micromolar range in enzyme- and/or cell-based assays and with high therapeutic indices. We also report the high-resolution X-ray cocrystal structures of NV 3CLpro-, poliovirus 3Cpro-, and transmissible gastroenteritis virus 3CLpro- GC376 inhibitor complexes, which show the compound covalently bound to a nucleophilic Cys residue in the catalytic site of the corresponding protease. We conclude that these compounds have the potential to be developed as antiviral therapeutics aimed at a single virus or multiple viruses in the picornavirus-like supercluster by targeting 3Cpro or 3CLpro.
Figures



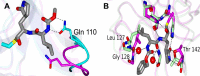

References
-
- Anand K, Ziebuhr J, Wadhwani P, Mesters JR, Hilgenfeld R. 2003. Coronavirus main proteinase (3CLpro) structure: basis for design of anti-SARS drugs. Science 300:1763–1767 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

