Regulated interaction of tegument proteins UL16 and UL11 from herpes simplex virus
- PMID: 22915809
- PMCID: PMC3486286
- DOI: 10.1128/JVI.01879-12
Regulated interaction of tegument proteins UL16 and UL11 from herpes simplex virus
Abstract
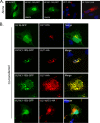
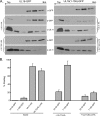
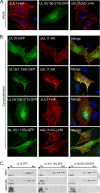
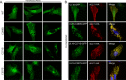
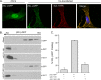
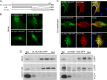
It is well known that proteins in the tegument (located between the viral capsid and envelope proteins) play critical roles in the assembly and budding of herpesviruses. Tegument proteins UL16 and UL11 of herpes simplex virus (HSV) are conserved among all the Herpesviridae. Although these proteins directly interact in vitro, UL16 was found to colocalize poorly with UL11 in cotransfected cells. To explain this discrepancy, we hypothesized that UL16 is initially made in an inactive form and is artificially transformed to the binding-competent state when cells are disrupted. Consistent with a regulated interaction, UL16 was able to fully colocalize with UL11 when a large C-terminal segment of UL16 was removed, creating mutant UL16(1-155). Moreover, membrane flotation assays revealed a massive movement of this mutant to the top of sucrose gradients in the presence of UL11, whereas both the full-length UL16 and the C-terminal fragment (residues 156 to 373) remained at the bottom. Further evidence for the presence of a C-terminal regulatory domain was provided by single-amino-acid substitutions at conserved cysteines (C269S, C271S, and C357S), which enabled the efficient interaction of full-length UL16 with UL11. Lastly, the binding site for UL11 was further mapped to residues 81 to 155, and to our surprise, the 5 Cys residues within UL16(1-155) are not required, even though the modification of free cysteines in UL16 with N-ethylmaleimide does in fact prevent binding. Collectively, these results reveal a regulatory function within the C-terminal region of UL16 that controls an N-terminal UL11-binding activity.
Figures

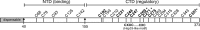

Similar articles
-
Analysis of the interaction between the UL11 and UL16 tegument proteins of herpes simplex virus.J Virol. 2008 Nov;82(21):10693-700. doi: 10.1128/JVI.01230-08. Epub 2008 Aug 20. J Virol. 2008. PMID: 18715918 Free PMC article.
-
Interaction domains of the UL16 and UL21 tegument proteins of herpes simplex virus.J Virol. 2010 Mar;84(6):2963-71. doi: 10.1128/JVI.02015-09. Epub 2009 Dec 30. J Virol. 2010. PMID: 20042500 Free PMC article.
-
Packaging determinants in the UL11 tegument protein of herpes simplex virus type 1.J Virol. 2006 Nov;80(21):10534-41. doi: 10.1128/JVI.01172-06. Epub 2006 Aug 23. J Virol. 2006. PMID: 16928743 Free PMC article.
-
Conserved tegument protein complexes: Essential components in the assembly of herpesviruses.Virus Res. 2015 Dec 2;210:308-17. doi: 10.1016/j.virusres.2015.09.007. Epub 2015 Sep 10. Virus Res. 2015. PMID: 26365681 Review.
-
Features and Functions of the Conserved Herpesvirus Tegument Protein UL11 and Its Binding Partners.Front Microbiol. 2022 Jun 3;13:829754. doi: 10.3389/fmicb.2022.829754. eCollection 2022. Front Microbiol. 2022. PMID: 35722336 Free PMC article. Review.
Cited by
-
Assembly and Egress of an Alphaherpesvirus Clockwork.Adv Anat Embryol Cell Biol. 2017;223:171-193. doi: 10.1007/978-3-319-53168-7_8. Adv Anat Embryol Cell Biol. 2017. PMID: 28528444 Free PMC article. Review.
-
Function of glycoprotein E of herpes simplex virus requires coordinated assembly of three tegument proteins on its cytoplasmic tail.Proc Natl Acad Sci U S A. 2012 Nov 27;109(48):19798-803. doi: 10.1073/pnas.1212900109. Epub 2012 Nov 12. Proc Natl Acad Sci U S A. 2012. PMID: 23150560 Free PMC article.
-
Duck enteritis virus UL21 is a late gene encoding a protein that interacts with pUL16.BMC Vet Res. 2020 Jan 8;16(1):8. doi: 10.1186/s12917-019-2228-7. BMC Vet Res. 2020. PMID: 31915010 Free PMC article.
-
The loss of both pUL16 and pUL21 in HSV-1-infected cells alters capsid-tegument composition, nuclear membrane architecture, cytoplasmic maturation and cell-to-cell spread.J Gen Virol. 2025 Mar;106(3):002083. doi: 10.1099/jgv.0.002083. J Gen Virol. 2025. PMID: 40080412 Free PMC article.
-
Long way up: rethink diseases in light of phase separation and phase transition.Protein Cell. 2024 Jul 1;15(7):475-492. doi: 10.1093/procel/pwad057. Protein Cell. 2024. PMID: 38069453 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

