Simple methods for generating and detecting locus-specific mutations induced with TALENs in the zebrafish genome
- PMID: 22916025
- PMCID: PMC3420959
- DOI: 10.1371/journal.pgen.1002861
Simple methods for generating and detecting locus-specific mutations induced with TALENs in the zebrafish genome
Abstract
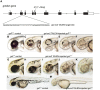
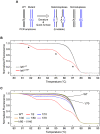
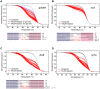
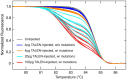
The zebrafish is a powerful experimental system for uncovering gene function in vertebrate organisms. Nevertheless, studies in the zebrafish have been limited by the approaches available for eliminating gene function. Here we present simple and efficient methods for inducing, detecting, and recovering mutations at virtually any locus in the zebrafish. Briefly, double-strand DNA breaks are induced at a locus of interest by synthetic nucleases, called TALENs. Subsequent host repair of the DNA lesions leads to the generation of insertion and deletion mutations at the targeted locus. To detect the induced DNA sequence alterations at targeted loci, genomes are examined using High Resolution Melt Analysis, an efficient and sensitive method for detecting the presence of newly arising sequence polymorphisms. As the DNA binding specificity of a TALEN is determined by a custom designed array of DNA recognition modules, each of which interacts with a single target nucleotide, TALENs with very high target sequence specificities can be easily generated. Using freely accessible reagents and Web-based software, and a very simple cloning strategy, a TALEN that uniquely recognizes a specific pre-determined locus in the zebrafish genome can be generated within days. Here we develop and test the activity of four TALENs directed at different target genes. Using the experimental approach described here, every embryo injected with RNA encoding a TALEN will acquire targeted mutations. Multiple independently arising mutations are produced in each growing embryo, and up to 50% of the host genomes may acquire a targeted mutation. Upon reaching adulthood, approximately 90% of these animals transmit targeted mutations to their progeny. Results presented here indicate the TALENs are highly sequence-specific and produce minimal off-target effects. In all, it takes about two weeks to create a target-specific TALEN and generate growing embryos that harbor an array of germ line mutations at a pre-specified locus.
Conflict of interest statement
DFV is a listed inventor on a patent application titled "TAL effector-mediated DNA modification" that is co-owned by Iowa State University and the University of Minnesota and that has been licensed to Cellectis, a European biotechnology company. All other authors have declared that no competing interests exist.
Figures
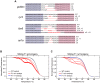

References
-
- Engert F, Wilson S (2011) Zebrafish neurobiology: From development to circuit function and behaviour. Dev Neurobiol - PubMed
-
- Hutson LD, Chien CB (2002) Wiring the zebrafish: axon guidance and synaptogenesis. Curr Opin Neurobiol 12: 87–92. - PubMed
-
- Krens SF, Heisenberg CP (2011) Cell sorting in development. Curr Top Dev Biol 95: 189–213. - PubMed
-
- Langdon YG, Mullins MC (2011) Maternal and zygotic control of zebrafish dorsoventral axial patterning. Annu Rev Genet 45: 357–377. - PubMed
-
- Lawson ND, Wolfe SA (2011) Forward and reverse genetic approaches for the analysis of vertebrate development in the zebrafish. Dev Cell 21: 48–64. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

