Identification of genetic modifiers of CagA-induced epithelial disruption in Drosophila
- PMID: 22919616
- PMCID: PMC3417398
- DOI: 10.3389/fcimb.2012.00024
Identification of genetic modifiers of CagA-induced epithelial disruption in Drosophila
Abstract
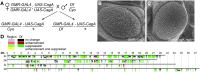
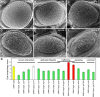
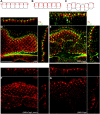
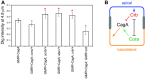
Helicobacter pylori strains containing the CagA protein are associated with high risk of gastric diseases including atrophic gastritis, peptic ulcers, and gastric cancer. CagA is injected into host cells via a Type IV secretion system where it activates growth factor-like signaling, disrupts cell-cell junctions, and perturbs host cell polarity. Using a transgenic Drosophila model, we have shown that CagA expression disrupts the morphogenesis of epithelial tissues such as the adult eye. Here we describe a genetic screen to identify modifiers of CagA-induced eye defects. We determined that reducing the copy number of genes encoding components of signaling pathways known to be targeted by CagA, such as the epidermal growth factor receptor (EGFR), modified the CagA-induced eye phenotypes. In our screen of just over half the Drosophila genome, we discovered 12 genes that either suppressed or enhanced CagA's disruption of the eye epithelium. Included in this list are genes involved in epithelial integrity, intracellular trafficking, and signal transduction. We investigated the mechanism of one suppressor, encoding the epithelial polarity determinant and junction protein Coracle, which is homologous to the mammalian Protein 4.1. We found that loss of a single copy of coracle improved the organization and integrity of larval retinal epithelia expressing CagA, but did not alter CagA's localization to cell junctions. Loss of a single copy of the coracle antagonist crumbs enhanced CagA-associated disruption of the larval retinal epithelium, whereas overexpression of crumbs suppressed this phenotype. Collectively, these results point to new cellular pathways whose disruption by CagA are likely to contribute to H. pylori-associated disease pathology.
Keywords: CagA; Drosophila; Helicobacter pylori; coracle; crumbs; epithelia; genetic modifier.
Figures

Similar articles
-
A transgenic Drosophila model demonstrates that the Helicobacter pylori CagA protein functions as a eukaryotic Gab adaptor.PLoS Pathog. 2008 May 16;4(5):e1000064. doi: 10.1371/journal.ppat.1000064. PLoS Pathog. 2008. PMID: 18483552 Free PMC article.
-
Helicobacter pylori stimulates epithelial cell migration via CagA-mediated perturbation of host cell signaling.Microbes Infect. 2012 May;14(5):470-6. doi: 10.1016/j.micinf.2011.12.003. Epub 2011 Dec 11. Microbes Infect. 2012. PMID: 22202178
-
CagA-ASPP2 complex mediates loss of cell polarity and favors H. pylori colonization of human gastric organoids.Proc Natl Acad Sci U S A. 2020 Feb 4;117(5):2645-2655. doi: 10.1073/pnas.1908787117. Epub 2020 Jan 21. Proc Natl Acad Sci U S A. 2020. PMID: 31964836 Free PMC article.
-
Influence of Helicobacter pylori virulence factors CagA and VacA on pathogenesis of gastrointestinal disorders.Microb Pathog. 2018 Apr;117:43-48. doi: 10.1016/j.micpath.2018.02.016. Epub 2018 Feb 9. Microb Pathog. 2018. PMID: 29432909 Review.
-
[Advances in CagA protein and CagAmediated pathogenesis of Helicobacter pylori-A review].Wei Sheng Wu Xue Bao. 2016 Dec 4;56(12):1821-30. Wei Sheng Wu Xue Bao. 2016. PMID: 29741846 Review. Chinese.
Cited by
-
Admixture Mapping of Alzheimer's disease in Caribbean Hispanics identifies a new locus on 22q13.1.Mol Psychiatry. 2022 Jun;27(6):2813-2820. doi: 10.1038/s41380-022-01526-6. Epub 2022 Apr 1. Mol Psychiatry. 2022. PMID: 35365809 Free PMC article.
-
Four Chromosomal Type IV Secretion Systems in Helicobacter pylori: Composition, Structure and Function.Front Microbiol. 2020 Jul 10;11:1592. doi: 10.3389/fmicb.2020.01592. eCollection 2020. Front Microbiol. 2020. PMID: 32754140 Free PMC article. Review.
-
Human pathogenic bacteria, fungi, and viruses in Drosophila: disease modeling, lessons, and shortcomings.Virulence. 2014 Feb 15;5(2):253-69. doi: 10.4161/viru.27524. Epub 2014 Jan 7. Virulence. 2014. PMID: 24398387 Free PMC article. Review.
-
Transgenic expression of the Helicobacter pylori virulence factor CagA promotes apoptosis or tumorigenesis through JNK activation in Drosophila.PLoS Pathog. 2012;8(10):e1002939. doi: 10.1371/journal.ppat.1002939. Epub 2012 Oct 18. PLoS Pathog. 2012. PMID: 23093933 Free PMC article.
-
Tight junction disruption: Helicobacter pylori and dysregulation of the gastric mucosal barrier.World J Gastroenterol. 2015 Oct 28;21(40):11411-27. doi: 10.3748/wjg.v21.i40.11411. World J Gastroenterol. 2015. PMID: 26523106 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

