Discovery of novel inhibitors of a disintegrin and metalloprotease 17 (ADAM17) using glycosylated and non-glycosylated substrates
- PMID: 22927435
- PMCID: PMC3476313
- DOI: 10.1074/jbc.M112.389114
Discovery of novel inhibitors of a disintegrin and metalloprotease 17 (ADAM17) using glycosylated and non-glycosylated substrates
Abstract
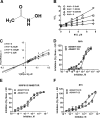
A disintegrin and metalloprotease (ADAM) proteases are implicated in multiple diseases, but no drugs based on ADAM inhibition exist. Most of the ADAM inhibitors developed to date feature zinc-binding moieties that target the active site zinc, which leads to a lack of selectivity and off-target toxicity. We hypothesized that secondary binding site (exosite) inhibitors should provide a viable alternative to active site inhibitors. Potential exosites in ADAM structures have been reported, but no studies describing substrate features necessary for exosite interactions exist. Analysis of ADAM cognate substrates revealed that glycosylation is often present in the vicinity of the scissile bond. To study whether glycosylation plays a role in modulating ADAM activity, a tumor necrosis factor α (TNFα) substrate with and without a glycan moiety attached was synthesized and characterized. Glycosylation enhanced ADAM8 and -17 activities and decreased ADAM10 activity. Metalloprotease (MMP) activity was unaffected by TNFα substrate glycosylation. High throughput screening assays were developed using glycosylated and non-glycosylated substrate, and positional scanning was conducted. A novel chemotype of ADAM17-selective probes was discovered from the TPIMS library (Houghten, R. A., Pinilla, C., Giulianotti, M. A., Appel, J. R., Dooley, C. T., Nefzi, A., Ostresh, J. M., Yu, Y., Maggiora, G. M., Medina-Franco, J. L., Brunner, D., and Schneider, J. (2008) Strategies for the use of mixture-based synthetic combinatorial libraries. Scaffold ranking, direct testing in vivo, and enhanced deconvolution by computational methods. J. Comb. Chem. 10, 3-19; Pinilla, C., Appel, J. R., Borràs, E., and Houghten, R. A. (2003) Advances in the use of synthetic combinatorial chemistry. Mixture-based libraries. Nat. Med. 9, 118-122) that preferentially inhibited glycosylated substrate hydrolysis and spared ADAM10, MMP-8, and MMP-14. Kinetic studies revealed that ADAM17 inhibition occurred via a non-zinc-binding mechanism. Thus, modulation of proteolysis via glycosylation may be used for identifying novel, potentially exosite binding compounds. The newly described ADAM17 inhibitors represent research tools to investigate the role of ADAM17 in the progression of various diseases.
Figures


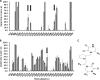
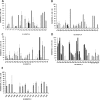

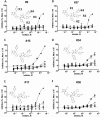
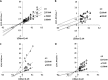
Similar articles
-
Activity of ADAM17 (a disintegrin and metalloprotease 17) is regulated by its noncatalytic domains and secondary structure of its substrates.J Biol Chem. 2013 Aug 2;288(31):22871-9. doi: 10.1074/jbc.M113.462267. Epub 2013 Jun 18. J Biol Chem. 2013. PMID: 23779109 Free PMC article.
-
Glycosylation of a disintegrin and metalloprotease 17 affects its activity and inhibition.Anal Biochem. 2014 Mar 15;449:68-75. doi: 10.1016/j.ab.2013.12.018. Epub 2013 Dec 19. Anal Biochem. 2014. PMID: 24361716 Free PMC article.
-
ADAMs 10 and 17 represent differentially regulated components of a general shedding machinery for membrane proteins such as transforming growth factor alpha, L-selectin, and tumor necrosis factor alpha.Mol Biol Cell. 2009 Mar;20(6):1785-94. doi: 10.1091/mbc.e08-11-1135. Epub 2009 Jan 21. Mol Biol Cell. 2009. PMID: 19158376 Free PMC article.
-
The "A Disintegrin And Metalloproteases" ADAM10 and ADAM17: novel drug targets with therapeutic potential?Eur J Cell Biol. 2011 Jun-Jul;90(6-7):527-35. doi: 10.1016/j.ejcb.2010.11.005. Epub 2010 Dec 30. Eur J Cell Biol. 2011. PMID: 21194787 Review.
-
The good, the bad and the ugly substrates for ADAM10 and ADAM17 in brain pathology, inflammation and cancer.Semin Cell Dev Biol. 2009 Apr;20(2):164-74. doi: 10.1016/j.semcdb.2008.09.005. Epub 2008 Sep 18. Semin Cell Dev Biol. 2009. PMID: 18951988 Review.
Cited by
-
Novel pyrrolidine diketopiperazines selectively inhibit melanoma cells via induction of late-onset apoptosis.J Med Chem. 2014 Feb 27;57(4):1599-608. doi: 10.1021/jm4019542. Epub 2014 Feb 5. J Med Chem. 2014. PMID: 24471466 Free PMC article.
-
Novel Approaches and Challenges of Discovery of Exosite Modulators of a Disintegrin and Metalloprotease 10.Front Mol Biosci. 2020 May 6;7:75. doi: 10.3389/fmolb.2020.00075. eCollection 2020. Front Mol Biosci. 2020. PMID: 32435655 Free PMC article. Review.
-
Identification of Bis-Cyclic Guanidines as Antiplasmodial Compounds from Positional Scanning Mixture-Based Libraries.Molecules. 2019 Mar 20;24(6):1100. doi: 10.3390/molecules24061100. Molecules. 2019. PMID: 30897744 Free PMC article.
-
Effects of Glycosylation on the Enzymatic Activity and Mechanisms of Proteases.Int J Mol Sci. 2016 Nov 25;17(12):1969. doi: 10.3390/ijms17121969. Int J Mol Sci. 2016. PMID: 27898009 Free PMC article. Review.
-
A Flow-Extension Tethered Particle Motion Assay for Single-Molecule Proteolysis.Biochemistry. 2019 May 21;58(20):2509-2518. doi: 10.1021/acs.biochem.9b00106. Epub 2019 Apr 12. Biochemistry. 2019. PMID: 30946563 Free PMC article.
References
-
- Houghten R. A., Pinilla C., Giulianotti M. A., Appel J. R., Dooley C. T., Nefzi A., Ostresh J. M., Yu Y., Maggiora G. M., Medina-Franco J. L., Brunner D., Schneider J. (2008) Strategies for the use of mixture-based synthetic combinatorial libraries. Scaffold ranking, direct testing in vivo, and enhanced deconvolution by computational methods. J. Comb. Chem. 10, 3–19 - PubMed
-
- Pinilla C., Appel J. R., Borràs E., Houghten R. A. (2003) Advances in the use of synthetic combinatorial chemistry. Mixture-based libraries. Nat. Med. 9, 118–122 - PubMed
-
- Moss M. L., Stoeck A., Yan W., Dempsey P. J. (2008) ADAM10 as a target for anti-cancer therapy. Curr. Pharm. Biotechnol. 9, 2–8 - PubMed
-
- Kataoka H. (2009) EGFR ligands and their signaling scissors, ADAMs, as new molecular targets for anticancer treatments. J. Dermatol. Sci. 56, 148–153 - PubMed
-
- Moss M. L., Sklair-Tavron L., Nudelman R. (2008) Drug insight. Tumor necrosis factor-converting enzyme as a pharmaceutical target for rheumatoid arthritis. Nat. Clin. Pract. Rheumatol. 4, 300–309 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

