Transcriptional profiling of long non-coding RNAs and novel transcribed regions across a diverse panel of archived human cancers
- PMID: 22929540
- PMCID: PMC4053743
- DOI: 10.1186/gb-2012-13-8-r75
Transcriptional profiling of long non-coding RNAs and novel transcribed regions across a diverse panel of archived human cancers
Abstract
Background: Molecular characterization of tumors has been critical for identifying important genes in cancer biology and for improving tumor classification and diagnosis. Long non-coding RNAs, as a new, relatively unstudied class of transcripts, provide a rich opportunity to identify both functional drivers and cancer-type-specific biomarkers. However, despite the potential importance of long non-coding RNAs to the cancer field, no comprehensive survey of long non-coding RNA expression across various cancers has been reported.
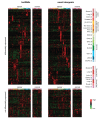
Results: We performed a sequencing-based transcriptional survey of both known long non-coding RNAs and novel intergenic transcripts across a panel of 64 archival tumor samples comprising 17 diagnostic subtypes of adenocarcinomas, squamous cell carcinomas and sarcomas. We identified hundreds of transcripts from among the known 1,065 long non-coding RNAs surveyed that showed variability in transcript levels between the tumor types and are therefore potential biomarker candidates. We discovered 1,071 novel intergenic transcribed regions and demonstrate that these show similar patterns of variability between tumor types. We found that many of these differentially expressed cancer transcripts are also expressed in normal tissues. One such novel transcript specifically expressed in breast tissue was further evaluated using RNA in situ hybridization on a panel of breast tumors. It was shown to correlate with low tumor grade and estrogen receptor expression, thereby representing a potentially important new breast cancer biomarker.
Conclusions: This study provides the first large survey of long non-coding RNA expression within a panel of solid cancers and also identifies a number of novel transcribed regions differentially expressed across distinct cancer types that represent candidate biomarkers for future research.
Figures



Similar articles
-
Long non-coding RNAs differentially expressed between normal versus primary breast tumor tissues disclose converse changes to breast cancer-related protein-coding genes.PLoS One. 2014 Sep 29;9(9):e106076. doi: 10.1371/journal.pone.0106076. eCollection 2014. PLoS One. 2014. PMID: 25264628 Free PMC article.
-
Pan-Cancer Analyses Reveal Long Intergenic Non-Coding RNAs Relevant to Tumor Diagnosis, Subtyping and Prognosis.EBioMedicine. 2016 May;7:62-72. doi: 10.1016/j.ebiom.2016.03.023. Epub 2016 Mar 19. EBioMedicine. 2016. PMID: 27322459 Free PMC article.
-
Transcriptional profiling of long-intergenic noncoding RNAs in lung squamous cell carcinoma and its value in diagnosis and prognosis.Mol Genet Genomic Med. 2019 Dec;7(12):e994. doi: 10.1002/mgg3.994. Epub 2019 Oct 16. Mol Genet Genomic Med. 2019. PMID: 31617686 Free PMC article.
-
Current Status of Long Non-Coding RNAs in Human Breast Cancer.Int J Mol Sci. 2016 Sep 6;17(9):1485. doi: 10.3390/ijms17091485. Int J Mol Sci. 2016. PMID: 27608009 Free PMC article. Review.
-
Long non-coding RNAs differential expression in breast cancer subtypes: What do we know?Clin Genet. 2019 May;95(5):558-568. doi: 10.1111/cge.13502. Epub 2019 Jan 22. Clin Genet. 2019. PMID: 30614523 Review.
Cited by
-
Discovery and characterization of long intergenic non-coding RNAs (lincRNA) module biomarkers in prostate cancer: an integrative analysis of RNA-Seq data.BMC Genomics. 2015;16 Suppl 7(Suppl 7):S3. doi: 10.1186/1471-2164-16-S7-S3. Epub 2015 Jun 11. BMC Genomics. 2015. PMID: 26100580 Free PMC article.
-
Genome-wide atlas of alternative polyadenylation in the forage legume red clover.Sci Rep. 2018 Jul 27;8(1):11379. doi: 10.1038/s41598-018-29699-7. Sci Rep. 2018. PMID: 30054540 Free PMC article.
-
HOXA10‑AS: A novel oncogenic long non‑coding RNA in glioma.Oncol Rep. 2018 Nov;40(5):2573-2583. doi: 10.3892/or.2018.6662. Epub 2018 Aug 21. Oncol Rep. 2018. PMID: 30132568 Free PMC article.
-
Polymorphisms in Long Noncoding RNA-Prostate Cancer-Associated Transcript 1 Are Associated with Lung Cancer Susceptibility in a Northeastern Chinese Population.DNA Cell Biol. 2019 Nov;38(11):1357-1365. doi: 10.1089/dna.2019.4834. Epub 2019 Aug 29. DNA Cell Biol. 2019. PMID: 31464517 Free PMC article.
-
Linc00675 is a novel marker of short survival and recurrence in patients with pancreatic ductal adenocarcinoma.World J Gastroenterol. 2015 Aug 21;21(31):9348-57. doi: 10.3748/wjg.v21.i31.9348. World J Gastroenterol. 2015. PMID: 26309360 Free PMC article.
References
-
- Perou CM, Sorlie T, Eisen MB, van de Rijn M, Jeffrey SS, Rees CA, Pollack JR, Ross DT, Johnsen H, Akslen LA, Fluge O, Pergamenschikov A, Williams C, Zhu SX, Lonning PE, Borresen-Dale AL, Brown PO, Botstein D. Molecular portraits of human breast tumours. Nature. 2000;13:747–752. doi: 10.1038/35021093. - DOI - PubMed
-
- Alizadeh AA, Eisen MB, Davis RE, Ma C, Lossos IS, Rosenwald A, Boldrick JC, Sabet H, Tran T, Yu X, Powell JI, Yang L, Marti GE, Moore T, Hudson J Jr, Lu L, Lewis DB, Tibshirani R, Sherlock G, Chan WC, Greiner TC, Weisenburger DD, Armitage JO, Warnke R, Levy R, Wilson W, Grever MR, Byrd JC, Botstein D, Brown PO. et al.Distinct types of diffuse large B-cell lymphoma identified by gene expression profiling. Nature. 2000;13:503–511. doi: 10.1038/35000501. - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

