Resolving conflict in eutherian mammal phylogeny using phylogenomics and the multispecies coalescent model
- PMID: 22930817
- PMCID: PMC3443116
- DOI: 10.1073/pnas.1211733109
Resolving conflict in eutherian mammal phylogeny using phylogenomics and the multispecies coalescent model
Erratum in
-
Correction for Song et al., Resolving conflict in eutherian mammal phylogeny using phylogenomics and the multispecies coalescent model.Proc Natl Acad Sci U S A. 2015 Nov 3;112(44):E6079. doi: 10.1073/pnas.1518753112. Epub 2015 Oct 26. Proc Natl Acad Sci U S A. 2015. PMID: 26504207 Free PMC article. No abstract available.
Abstract
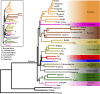
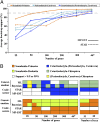
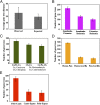
The reconstruction of the Tree of Life has relied almost entirely on concatenation methods, which do not accommodate gene tree heterogeneity, a property that simulations and theory have identified as a likely cause of incongruent phylogenies. However, this incongruence has not yet been demonstrated in empirical studies. Several key relationships among eutherian mammals remain controversial and conflicting among previous studies, including the root of eutherian tree and the relationships within Euarchontoglires and Laurasiatheria. Both bayesian and maximum-likelihood analysis of genome-wide data of 447 nuclear genes from 37 species show that concatenation methods indeed yield strong incongruence in the phylogeny of eutherian mammals, as revealed by subsampling analyses of loci and taxa, which produced strongly conflicting topologies. In contrast, the coalescent methods, which accommodate gene tree heterogeneity, yield a phylogeny that is robust to variable gene and taxon sampling and is congruent with geographic data. The data also demonstrate that incomplete lineage sorting, a major source of gene tree heterogeneity, is relevant to deep-level phylogenies, such as those among eutherian mammals. Our results firmly place the eutherian root between Atlantogenata and Boreoeutheria and support ungulate polyphyly and a sister-group relationship between Scandentia and Primates. This study demonstrates that the incongruence introduced by concatenation methods is a major cause of long-standing uncertainty in the phylogeny of eutherian mammals, and the same may apply to other clades. Our analyses suggest that such incongruence can be resolved using phylogenomic data and coalescent methods that deal explicitly with gene tree heterogeneity.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
Comment in
-
Concatenation versus coalescence versus "concatalescence".Proc Natl Acad Sci U S A. 2013 Mar 26;110(13):E1179. doi: 10.1073/pnas.1221121110. Epub 2013 Mar 4. Proc Natl Acad Sci U S A. 2013. PMID: 23460702 Free PMC article. No abstract available.
-
Reply to Gatesy and Springer: the multispecies coalescent model can effectively handle recombination and gene tree heterogeneity.Proc Natl Acad Sci U S A. 2013 Mar 26;110(13):E1180. doi: 10.1073/pnas.1300129110. Proc Natl Acad Sci U S A. 2013. PMID: 23650651 Free PMC article. No abstract available.
References
-
- de Queiroz A, Gatesy J. The supermatrix approach to systematics. Trends Ecol Evol. 2007;22(1):34–41. - PubMed
-
- William J, Ballard O. Combining data in phylogenetic analysis. Trends Ecol Evol. 1996;11:334. - PubMed
-
- Belfiore NM, Liu L, Moritz C. Multilocus phylogenetics of a rapid radiation in the genus Thomomys (Rodentia: Geomyidae) Syst Biol. 2008;57(2):294–310. - PubMed
-
- Edwards SV. Is a new and general theory of molecular systematics emerging? Evolution. 2009;63(1):1–19. - PubMed
-
- Kubatko LS, Degnan JH. Inconsistency of phylogenetic estimates from concatenated data under coalescence. Syst Biol. 2007;56(1):17–24. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

