Specific genomic regions are differentially affected by copy number alterations across distinct cancer types, in aggregated cytogenetic data
- PMID: 22937079
- PMCID: PMC3427184
- DOI: 10.1371/journal.pone.0043689
Specific genomic regions are differentially affected by copy number alterations across distinct cancer types, in aggregated cytogenetic data
Abstract
Background: Regional genomic copy number alterations (CNA) are observed in the vast majority of cancers. Besides specifically targeting well-known, canonical oncogenes, CNAs may also play more subtle roles in terms of modulating genetic potential and broad gene expression patterns of developing tumors. Any significant differences in the overall CNA patterns between different cancer types may thus point towards specific biological mechanisms acting in those cancers. In addition, differences among CNA profiles may prove valuable for cancer classifications beyond existing annotation systems.
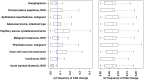
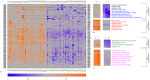
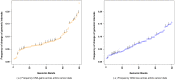
Principal findings: We have analyzed molecular-cytogenetic data from 25579 tumors samples, which were classified into 160 cancer types according to the International Classification of Disease (ICD) coding system. When correcting for differences in the overall CNA frequencies between cancer types, related cancers were often found to cluster together according to similarities in their CNA profiles. Based on a randomization approach, distance measures from the cluster dendrograms were used to identify those specific genomic regions that contributed significantly to this signal. This approach identified 43 non-neutral genomic regions whose propensity for the occurrence of copy number alterations varied with the type of cancer at hand. Only a subset of these identified loci overlapped with previously implied, highly recurrent (hot-spot) cytogenetic imbalance regions.
Conclusions: Thus, for many genomic regions, a simple null-hypothesis of independence between cancer type and relative copy number alteration frequency can be rejected. Since a subset of these regions display relatively low overall CNA frequencies, they may point towards second-tier genomic targets that are adaptively relevant but not necessarily essential for cancer development.
Conflict of interest statement
Figures


Similar articles
-
CNApp, a tool for the quantification of copy number alterations and integrative analysis revealing clinical implications.Elife. 2020 Jan 15;9:e50267. doi: 10.7554/eLife.50267. Elife. 2020. PMID: 31939734 Free PMC article.
-
Landscape of somatic allelic imbalances and copy number alterations in HER2-amplified breast cancer.Breast Cancer Res. 2011;13(6):R129. doi: 10.1186/bcr3075. Epub 2011 Dec 14. Breast Cancer Res. 2011. PMID: 22169037 Free PMC article.
-
Recurrent patterns of DNA copy number alterations in tumors reflect metabolic selection pressures.Mol Syst Biol. 2017 Feb 15;13(2):914. doi: 10.15252/msb.20167159. Mol Syst Biol. 2017. PMID: 28202506 Free PMC article.
-
Interpretation of copy number alterations identified through clinical microarray-comparative genomic hybridization.Clin Lab Med. 2011 Dec;31(4):565-80, viii. doi: 10.1016/j.cll.2011.08.007. Clin Lab Med. 2011. PMID: 22118737 Review.
-
Copy number alteration signatures as biomarkers in cancer: a review.Biomark Med. 2022 Apr;16(5):371-386. doi: 10.2217/bmm-2021-0476. Epub 2022 Feb 23. Biomark Med. 2022. PMID: 35195030 Review.
Cited by
-
Candidate targets of copy number deletion events across 17 cancer types.Front Genet. 2023 Jan 16;13:1017657. doi: 10.3389/fgene.2022.1017657. eCollection 2022. Front Genet. 2023. PMID: 36726722 Free PMC article.
-
arrayMap 2014: an updated cancer genome resource.Nucleic Acids Res. 2015 Jan;43(Database issue):D825-30. doi: 10.1093/nar/gku1123. Epub 2014 Nov 26. Nucleic Acids Res. 2015. PMID: 25428357 Free PMC article.
References
-
- Kallioniemi A, Kallioniemi OP, Sudar D, Rutovitz D, Gray JW, et al. (1992) Comparative genomic hybridization for molecular cytogenetic analysis of solid tumors. Science 258: 818–21. - PubMed
-
- Joos S, Scherthan H, Speicher MR, Schlegel J, Cremer T, et al. (1993) Detection of amplified dna sequences by reverse chromosome painting using genomic tumor dna as probe. Hum Genet 90: 584–9. - PubMed
-
- Solinas-Toldo S, Lampel S, Stilgenbauer S, Nickolenko J, Benner A, et al. (1997) Matrix-based comparative genomic hybridization: biochips to screen for genomic imbalances. Genes Chromosomes Cancer 20: 399–407. - PubMed
-
- Pinkel D, Segraves R, Sudar D, Clark S, Poole I, et al. (1998) High resolution analysis of dna copy number variation using comparative genomic hybridization to microarrays. Nat Genet 20: 207–11. - PubMed
MeSH terms
LinkOut - more resources
Full Text Sources

