Measures of compositional strand bias related to replication machinery and its applications
- PMID: 22942671
- PMCID: PMC3269016
- DOI: 10.2174/138920212799034749
Measures of compositional strand bias related to replication machinery and its applications
Abstract
The compositional asymmetry of complementary bases in nucleotide sequences implies the existence of a mutational or selectional bias in the two strands of the DNA duplex, which is commonly shaped by strand-specific mechanisms in transcription or replication. Such strand bias in genomes, frequently visualized by GC skew graphs, is used for the computational prediction of transcription start sites and replication origins, as well as for comparative evolutionary genomics studies. The use of measures of compositional strand bias in order to quantify the degree of strand asymmetry is crucial, as it is the basis for determining the applicability of compositional analysis and comparing the strength of the mutational bias in different biological machineries in various species. Here, we review the measures of strand bias that have been proposed to date, including the ∆GC skew, the B(1) index, the predictability score of linear discriminant analysis for gene orientation, the signal-to-noise ratio of the oligonucleotide bias, and the GC skew index. These measures have been predominantly designed for and applied to the analysis of replication-related mutational processes in prokaryotes, but we also give research examples in eukaryotes.
Keywords: GC skew; Nucleotide composition bias; bacterial replication; replication-related mutations..
Figures
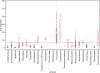

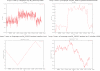
References
-
- Nakabachi A, Yamashita A, Toh H, Ishikawa H, Dunbar H E, Moran N A, Hattori M. The 160-kilobase genome of the bacterial endosymbiont Carsonella. Science. 2006;314(5797 ):267. - PubMed
-
- Rocha E P, Danchin A. Base composition bias might result from competition for metabolic resources. Trends Genet. 2002;18(6 ):291–294. - PubMed
-
- Mitchell D. GC content and genome length in Chargaff compliant genomes. Biochem. Biophys. Res. Commun. 2007;353(1 ):207–210. - PubMed
LinkOut - more resources
Full Text Sources
Miscellaneous
