Analysis of T-DNA alleles of flavonoid biosynthesis genes in Arabidopsis ecotype Columbia
- PMID: 22947320
- PMCID: PMC3526476
- DOI: 10.1186/1756-0500-5-485
Analysis of T-DNA alleles of flavonoid biosynthesis genes in Arabidopsis ecotype Columbia
Erratum in
-
Correction: Analysis of T-DNA alleles of flavonoid biosynthesis genes in Arabidopsis ecotype Columbia.BMC Res Notes. 2025 Jul 31;18(1):338. doi: 10.1186/s13104-025-07316-x. BMC Res Notes. 2025. PMID: 40745661 Free PMC article. No abstract available.
Abstract
Background: The flavonoid pathway is a long-standing and important tool for plant genetics, biochemistry, and molecular biology. Numerous flavonoid mutants have been identified in Arabidopsis over the past several decades in a variety of ecotypes. Here we present an analysis of Arabidopsis lines of ecotype Columbia carrying T-DNA insertions in genes encoding enzymes of the central flavonoid pathway. We also provide a comprehensive summary of various mutant alleles for these structural genes that have been described in the literature to date in a wide variety of ecotypes.
Findings: The confirmed knockout lines present easily-scorable phenotypes due to altered pigmentation of the seed coat (or testa). Knockouts for seven alleles for six flavonoid biosynthetic genes were confirmed by PCR and characterized by UPLC for altered flavonol content.
Conclusion: Seven mutant lines for six genes of the central flavonoid pathway were characterized in ecotype, Columbia. These lines represent a useful resource for integrating biochemical and physiological studies with genomic, transcriptomic, and proteomic data, much of which has been, and continues to be, generated in the Columbia background.
Figures



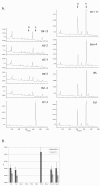
References
-
- Winkel BSJ. In: The Science of Flavonoids. Grotewold E, editor. New York: Springer Science & Business Media; 2006. The biosynthesis of flavonoids; pp. 71–95.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

