MAPK15/ERK8 stimulates autophagy by interacting with LC3 and GABARAP proteins
- PMID: 22948227
- PMCID: PMC3541284
- DOI: 10.4161/auto.21857
MAPK15/ERK8 stimulates autophagy by interacting with LC3 and GABARAP proteins
Abstract
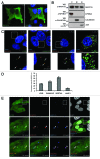
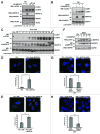
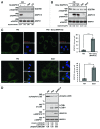
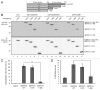
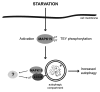
Macroautophagy (hereafter referred to as autophagy) is an evolutionarily conserved catabolic process necessary for normal recycling of cellular constituents and for appropriate response to cellular stress. Although several genes belonging to the core molecular machinery involved in autophagosome formation have been discovered, relatively little is known about the nature of signaling networks controlling autophagy upon intracellular or extracellular stimuli. We discovered ATG8-like proteins (MAP1LC3B, GABARAP and GABARAPL1) as novel interactors of MAPK15/ERK8, a MAP kinase involved in cell proliferation and transformation. Based on the role of these proteins in the autophagic process, we demonstrated that MAPK15 is indeed localized to autophagic compartments and increased, in a kinase-dependent fashion, ATG8-like proteins lipidation, autophagosome formation and SQSTM1 degradation, while decreasing LC3B inhibitory phosphorylation. Interestingly, we also identified a conserved LC3-interacting region (LIR) in MAPK15 responsible for its interaction with ATG8-like proteins, for its localization to autophagic structures and, consequently, for stimulation of the formation of these compartments. Furthermore, we reveal that MAPK15 activity was induced in response to serum and amino-acid starvation and that this stimulus, in turn, required endogenous MAPK15 expression to induce the autophagic process. Altogether, these results suggested a new function for MAPK15 as a regulator of autophagy, acting through interaction with ATG8 family proteins. Also, based on the key role of this process in several human diseases, these results supported the use of this MAP kinase as a potential novel therapeutic target.
Figures



Similar articles
-
Members of the autophagy class III phosphatidylinositol 3-kinase complex I interact with GABARAP and GABARAPL1 via LIR motifs.Autophagy. 2019 Aug;15(8):1333-1355. doi: 10.1080/15548627.2019.1581009. Epub 2019 Mar 4. Autophagy. 2019. PMID: 30767700 Free PMC article.
-
Redundancy of human ATG4 protease isoforms in autophagy and LC3/GABARAP processing revealed in cells.Autophagy. 2019 Jun;15(6):976-997. doi: 10.1080/15548627.2019.1569925. Epub 2019 Feb 1. Autophagy. 2019. PMID: 30661429 Free PMC article.
-
Development of LC3/GABARAP sensors containing a LIR and a hydrophobic domain to monitor autophagy.EMBO J. 2017 Apr 13;36(8):1100-1116. doi: 10.15252/embj.201696315. Epub 2017 Mar 20. EMBO J. 2017. PMID: 28320742 Free PMC article.
-
LC3/GABARAP family proteins: autophagy-(un)related functions.FASEB J. 2016 Dec;30(12):3961-3978. doi: 10.1096/fj.201600698R. Epub 2016 Sep 6. FASEB J. 2016. PMID: 27601442 Review.
-
A new perspective on the autophagic and non-autophagic functions of the GABARAP protein family: a potential therapeutic target for human diseases.Mol Cell Biochem. 2024 Jun;479(6):1415-1441. doi: 10.1007/s11010-023-04800-5. Epub 2023 Jul 13. Mol Cell Biochem. 2024. PMID: 37440122 Review.
Cited by
-
Disturbed Flow Induces Autophagy, but Impairs Autophagic Flux to Perturb Mitochondrial Homeostasis.Antioxid Redox Signal. 2015 Nov 20;23(15):1207-19. doi: 10.1089/ars.2014.5896. Epub 2015 Jun 29. Antioxid Redox Signal. 2015. PMID: 26120766 Free PMC article.
-
Autophagy processes are dependent on EGF receptor signaling.Oncotarget. 2018 Jul 13;9(54):30289-30303. doi: 10.18632/oncotarget.25708. eCollection 2018 Jul 13. Oncotarget. 2018. PMID: 30100990 Free PMC article.
-
MAPK15 protects from oxidative stress-dependent cellular senescence by inducing the mitophagic process.Aging Cell. 2022 Jul;21(7):e13620. doi: 10.1111/acel.13620. Epub 2022 Jun 1. Aging Cell. 2022. PMID: 35642724 Free PMC article.
-
Atypical MAP kinases - new insights and directions from amoeba.J Cell Sci. 2023 Oct 15;136(20):jcs261447. doi: 10.1242/jcs.261447. Epub 2023 Oct 16. J Cell Sci. 2023. PMID: 37850857 Free PMC article. Review.
-
Guidelines for the use and interpretation of assays for monitoring autophagy (3rd edition).Autophagy. 2016;12(1):1-222. doi: 10.1080/15548627.2015.1100356. Autophagy. 2016. PMID: 26799652 Free PMC article. No abstract available.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
