Identification of hepatitis C virus inhibitors targeting different aspects of infection using a cell-based assay
- PMID: 22948883
- PMCID: PMC3497158
- DOI: 10.1128/AAC.01413-12
Identification of hepatitis C virus inhibitors targeting different aspects of infection using a cell-based assay
Abstract
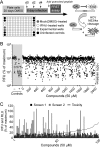
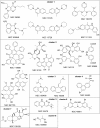
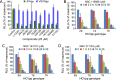
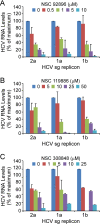
With 2 to 3% of the worldwide population chronically infected, hepatitis C virus (HCV) infection continues to be a major health care burden. Unfortunately, current interferon-based treatment options are not effective in all patients and are associated with significant side effects. Consequently, there is an ongoing need to identify and develop new anti-HCV therapies. Toward this goal, we previously developed a cell-based HCV infection assay for antiviral compound screening based on a low-multiplicity-of-infection approach that uniquely allows for the identification of antiviral compounds that target cell culture-derived HCV (HCVcc) at any step of the viral infection cycle. Using this assay, here we report the screening of the NCI Diversity Set II library, containing 1,974 synthesized chemical compounds, and the identification of compounds with specific anti-HCV activity. In combination with toxicity counterscreening, we identified 30 hits from the compound library, 13 of which showed reproducible and dose-dependent inhibition of HCV with mean therapeutic indices (50% cytotoxic concentration [CC(50)]/50% effective concentration [EC(50)]) of greater than 6. Using HCV pseudotype and replicon systems of multiple HCV genotypes, as well as infectious HCVcc-based assembly and secretion analysis, we determined that different compounds within this group of candidate inhibitors target different steps of viral infection. The compounds identified not only will serve as biological probes to study and further dissect the biology of viral infection but also should facilitate the development of new anti-HCV therapeutic treatments.
Figures

References
-
- Afdhal NH. 2004. The natural history of hepatitis C. Semin. Liver Dis. 24(Suppl 2):3–8 - PubMed
-
- Ahmed A, Keeffe EB. 1999. Treatment strategies for chronic hepatitis C: update since the 1997 National Institutes of Health Consensus Development Conference. J. Gastroenterol. Hepatol. 14(Suppl):S12–S18 - PubMed
-
- Ahmed-Belkacem A, et al. 2010. Silibinin and related compounds are direct inhibitors of hepatitis C virus RNA-dependent RNA polymerase. Gastroenterology 138:1112–1122 - PubMed
-
- Alter HJ, Seeff LB. 2000. Recovery, persistence, and sequelae in hepatitis C virus infection: a perspective on long-term outcome. Semin. Liver Dis. 20:17–35 - PubMed
-
- Armstrong GL, et al. 2006. The prevalence of hepatitis C virus infection in the United States, 1999 through 2002. Ann. Intern. Med. 144:705–714 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

