Improper tagging of the non-essential small capsid protein VP26 impairs nuclear capsid egress of herpes simplex virus
- PMID: 22952920
- PMCID: PMC3432071
- DOI: 10.1371/journal.pone.0044177
Improper tagging of the non-essential small capsid protein VP26 impairs nuclear capsid egress of herpes simplex virus
Abstract
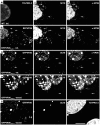
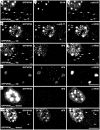
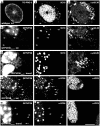
To analyze the subcellular trafficking of herpesvirus capsids, the small capsid protein has been labeled with different fluorescent proteins. Here, we analyzed the infectivity of several HSV1(17(+)) strains in which the N-terminal region of the non-essential small capsid protein VP26 had been tagged at different positions. While some variants replicated with similar kinetics as their parental wild type strain, others were not infectious at all. Improper tagging resulted in the aggregation of VP26 in the nucleus, prevented efficient nuclear egress of viral capsids, and thus virion formation. Correlative fluorescence and electron microscopy showed that these aggregates had sequestered several other viral proteins, but often did not contain viral capsids. The propensity for aggregate formation was influenced by the type of the fluorescent protein domain, the position of the inserted tag, the cell type, and the progression of infection. Among the tags that we have tested, mRFPVP26 had the lowest tendency to induce nuclear aggregates, and showed the least reduction in replication when compared to wild type. Our data suggest that bona fide monomeric fluorescent protein tags have less impact on proper assembly of HSV1 capsids and nuclear capsid egress than tags that tend to dimerize. Small chemical compounds capable of inducing aggregate formation of VP26 may lead to new antiviral drugs against HSV infections.
Conflict of interest statement
Figures




References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

