Tigecycline challenge triggers sRNA production in Salmonella enterica serovar Typhimurium
- PMID: 22958399
- PMCID: PMC3511261
- DOI: 10.1186/1471-2180-12-195
Tigecycline challenge triggers sRNA production in Salmonella enterica serovar Typhimurium
Abstract
Background: Bacteria employ complex transcriptional networks involving multiple genes in response to stress, which is not limited to gene and protein networks but now includes small RNAs (sRNAs). These regulatory RNA molecules are increasingly shown to be able to initiate regulatory cascades and modulate the expression of multiple genes that are involved in or required for survival under environmental challenge. Despite mounting evidence for the importance of sRNAs in stress response, their role upon antibiotic exposure remains unknown. In this study, we sought to determine firstly, whether differential expression of sRNAs occurs upon antibiotic exposure and secondly, whether these sRNAs could be attributed to microbial tolerance to antibiotics.
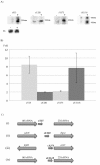
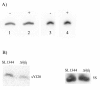
Results: A small scale sRNA cloning strategy of Salmonella enterica serovar Typhimurium SL1344 challenged with half the minimal inhibitory concentration of tigecycline identified four sRNAs (sYJ5, sYJ20, sYJ75 and sYJ118) which were reproducibly upregulated in the presence of either tigecycline or tetracycline. The coding sequences of the four sRNAs were found to be conserved across a number of species. Genome analysis found that sYJ5 and sYJ118 mapped between the 16S and 23S rRNA encoding genes. sYJ20 (also known as SroA) is encoded upstream of the tbpAyabKyabJ operon and is classed as a riboswitch, whilst its role in antibiotic stress-response appears independent of its riboswitch function. sYJ75 is encoded between genes that are involved in enterobactin transport and metabolism. Additionally we find that the genetic deletion of sYJ20 rendered a reduced viability phenotype in the presence of tigecycline, which was recovered when complemented. The upregulation of some of these sRNAs were also observed when S. Typhimurium was challenged by ampicillin (sYJ5, 75 and 118); or when Klebsiella pneumoniae was challenged by tigecycline (sYJ20 and 118).
Conclusions: Small RNAs are overexpressed as a result of antibiotic exposure in S. Typhimurium where the same molecules are upregulated in a related species or after exposure to different antibiotics. sYJ20, a riboswitch, appears to possess a trans-regulatory sRNA role in antibiotic tolerance. These findings imply that the sRNA mediated response is a component of the bacterial response to antibiotic challenge.
Figures


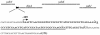


References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

