Overexpression of Arabidopsis plasmodesmata germin-like proteins disrupts root growth and development
- PMID: 22960910
- PMCID: PMC3480292
- DOI: 10.1105/tpc.112.101063
Overexpression of Arabidopsis plasmodesmata germin-like proteins disrupts root growth and development
Abstract
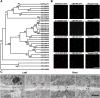
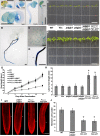
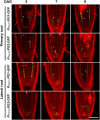
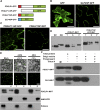
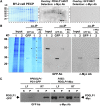
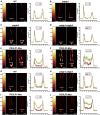
In plants, a population of non-cell-autonomous proteins (NCAPs), including numerous transcription factors, move cell to cell through plasmodesmata (PD). In many cases, the intercellular trafficking of these NCAPs is regulated by their interaction with specific PD components. To gain further insight into the functions of this NCAP pathway, coimmunoprecipitation experiments were performed on a tobacco (Nicotiana tabacum) plasmodesmal-enriched cell wall protein preparation using as bait the NCAP, pumpkin (Cucurbita maxima) PHLOEM PROTEIN16 (Cm-PP16). A Cm-PP16 interaction partner, Nt-PLASMODESMAL GERMIN-LIKE PROTEIN1 (Nt-PDGLP1) was identified and shown to be a PD-located component. Arabidopsis thaliana putative orthologs, PDGLP1 and PDGLP2, were identified; expression studies indicated that, postgermination, these proteins were preferentially expressed in the root system. The PDGLP1 signal peptide was shown to function in localization to the PD by a novel mechanism involving the endoplasmic reticulum-Golgi secretory pathway. Overexpression of various tagged versions altered root meristem function, leading to reduced primary root but enhanced lateral root growth. This effect on root growth was corrected with an inability of these chimeric proteins to form stable PD-localized complexes. PDGLP1 and PDGLP2 appear to be involved in regulating primary root growth by controlling phloem-mediated allocation of resources between the primary and lateral root meristems.
Figures



Similar articles
-
Reciprocal phosphorylation and glycosylation recognition motifs control NCAPP1 interaction with pumpkin phloem proteins and their cell-to-cell movement.Plant Cell. 2007 Jun;19(6):1866-84. doi: 10.1105/tpc.107.052522. Epub 2007 Jun 29. Plant Cell. 2007. PMID: 17601822 Free PMC article.
-
Early development and gravitropic response of lateral roots in Arabidopsis thaliana.Philos Trans R Soc Lond B Biol Sci. 2012 Jun 5;367(1595):1509-16. doi: 10.1098/rstb.2011.0231. Philos Trans R Soc Lond B Biol Sci. 2012. PMID: 22527393 Free PMC article.
-
Class 1 reversibly glycosylated polypeptides are plasmodesmal-associated proteins delivered to plasmodesmata via the golgi apparatus.Plant Cell. 2005 Jun;17(6):1788-800. doi: 10.1105/tpc.105.031823. Epub 2005 May 6. Plant Cell. 2005. PMID: 15879561 Free PMC article.
-
Transcription factors on the move.Curr Opin Plant Biol. 2012 Dec;15(6):645-51. doi: 10.1016/j.pbi.2012.09.010. Epub 2012 Sep 29. Curr Opin Plant Biol. 2012. PMID: 23031575 Review.
-
The role of mobile small RNA species during root growth and development.Curr Opin Cell Biol. 2012 Apr;24(2):211-6. doi: 10.1016/j.ceb.2011.12.005. Epub 2012 Jan 5. Curr Opin Cell Biol. 2012. PMID: 22227227 Review.
Cited by
-
Plasmodesmata dynamics are coordinated by intracellular signaling pathways.Curr Opin Plant Biol. 2013 Oct;16(5):614-20. doi: 10.1016/j.pbi.2013.07.007. Epub 2013 Aug 23. Curr Opin Plant Biol. 2013. PMID: 23978390 Free PMC article. Review.
-
Roles of Germin-like Protein Family in Response to Seed Germination and Shoot Branching in Brassica napus.Int J Mol Sci. 2024 Oct 26;25(21):11518. doi: 10.3390/ijms252111518. Int J Mol Sci. 2024. PMID: 39519071 Free PMC article.
-
The transcriptomics of an experimentally evolved plant-virus interaction.Sci Rep. 2016 Apr 26;6:24901. doi: 10.1038/srep24901. Sci Rep. 2016. PMID: 27113435 Free PMC article.
-
Plasmodesmata-mediated intercellular signaling during plant growth and development.Front Plant Sci. 2014 Feb 17;5:44. doi: 10.3389/fpls.2014.00044. eCollection 2014. Front Plant Sci. 2014. PMID: 24596574 Free PMC article.
-
Genome-Wide Characterization and Expression Analysis of the Germin-Like Protein Family in Rice and Arabidopsis.Int J Mol Sci. 2016 Sep 23;17(10):1622. doi: 10.3390/ijms17101622. Int J Mol Sci. 2016. PMID: 27669230 Free PMC article.
References
-
- Bernier F., Berna A. (2001). Germins and germin-like proteins: Plant do-all proteins. But what do they do exactly? Plant Physiol. Biochem. 39: 545–554
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

