Identification and characterization of five intramembrane metalloproteases in Anabaena variabilis
- PMID: 22961855
- PMCID: PMC3486370
- DOI: 10.1128/JB.01366-12
Identification and characterization of five intramembrane metalloproteases in Anabaena variabilis
Abstract
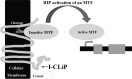
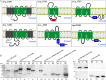
Regulated intramembrane proteolysis (RIP) involves cleavage of a transmembrane segment of a protein, releasing the active form of a membrane-anchored transcription factor (MTF) or a membrane-tethered signaling protein in response to an extracellular or intracellular signal. RIP is conserved from bacteria to humans and governs many important signaling pathways in both prokaryotes and eukaryotes. Proteases that carry out these cleavages are named intramembrane cleaving proteases (I-CLips). To date, little is known about I-CLips in cyanobacteria. In this study, five putative site-2 type I-Clips (Ava_1070, Ava_1730, Ava_1797, Ava_3438, and Ava_4785) were identified through a genome-wide survey in Anabaena variabilis. Biochemical analysis demonstrated that these five putative A. variabilis site-2 proteases (S2Ps(Av)) have authentic protease activities toward an artificial substrate pro-σ(K), a Bacillus subtilis MTF, in our reconstituted Escherichia coli system. The enzymatic activities of processing pro-σ(K) differ among these five S2Ps(Av). Substitution of glutamic acid (E) by glutamine (Q) in the conserved HEXXH zinc-coordinated motif caused the loss of protease activities in these five S2Ps(Av), suggesting that they belonged to the metalloprotease family. Further mapping of the cleaved peptides of pro-σ(K) by Ava_4785 and Ava_1797 revealed that Ava_4785 and Ava_1797 recognized the same cleavage site in pro-σ(K) as SpoIVFB, a cognate S2P of pro-σ(K) from B. subtilis. Taking these results together, we report here for the first time the identification of five metallo-intramembrane cleaving proteases in Anabaena variabilis. The experimental system described herein should be applicable to studies of other RIP events and amenable to developing in vitro assays for I-CLips.
Figures



References
-
- Adams DG, Duggan PS. 1999. Tansley review no. 107. Heterocyst and akinete differentiation in cyanobacteria. New Phytol. 144:3–33
-
- Ali N, Knaüper V. 2007. Phorbol ester-induced shedding of the prostate cancer marker transmembrane protein with epidermal growth factor and two follistatin motifs 2 is mediated by the disintegrin and metalloproteinase-17. J. Biol. Chem. 282:37378–37388 - PubMed
-
- Blot G, et al. 2006. Luman, a new partner of HIV-1 TMgp41, interferes with Tat-mediated transcription of the HIV-1 LTR. J. Mol. Biol. 364:1034–1047 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

