Molecular phylogeny, laboratory rearing, and karyotype of the bombycid moth, Trilocha varians
- PMID: 22963522
- PMCID: PMC3476958
- DOI: 10.1673/031.012.4901
Molecular phylogeny, laboratory rearing, and karyotype of the bombycid moth, Trilocha varians
Abstract
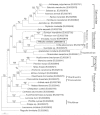
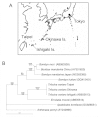
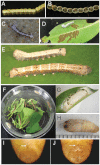
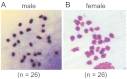
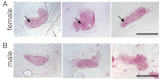
This study describes the molecular phylogeny, laboratory rearing, and karyotype of a bombycid moth, Trilocha varians (F. Walker) (Lepidoptera: Bombycidae), which feeds on leaves of Ficus spp. (Rosales: Moraceae). The larvae of this species were collected in Taipei city, Taiwan, and the Ryukyu Archipelago (Ishigaki and Okinawa Islands, Japan). Molecular phylogenetic analyses revealed that T. varians belongs to the subfamily Bombycinae, thus showing a close relationship to the domesticated silkworm Bombyx mori (L.), a lepidopteran model insect. A laboratory method was developed for rearing T. varians and the time required for development from the embryo to adult was determined. From oviposition to adult emergence, the developmental zero was 10.47 °C and total effective temperature was 531.2 day-degrees, i.e., approximately 30 days for one generation when reared at 28 °C. The haploid of T. varians consisted of n = 26 chromosomes. In highly polyploid somatic nuclei, females showed a large heterochromatin body, indicating that the sex chromosome system in T. varians is WZ/ZZ (female/male). The results of the present study should facilitate the utilization of T. varians as a reference species for B. mori, thereby leading to a greater understanding of the ecology and evolution of bombycid moths.
Figures

References
-
- Akaike H. A new look at the statistical model identification. IEEE Transactions on Automatic Control. 1974;19:716–723.
-
- Arunkumar KP, Metta M, Nagaraju J. Molecular phylogeny of silkmoths reveals the origin of domesticated silkmoth, Bombyx mori from Chinese Bombyx mandarina and paternal inheritance of Antheraea proylei mitochondrial DNA. Molecular Phylogenetics and Evolution. 2006;40:419–427. - PubMed
-
- Banno Y, Fujii H, Kawaguchi Y, Yamamoto K, Nishikawa K, Nishisaka A, Tamura K, Eguchi S. A Guide to the Silkworm Mutants 2005: Gene name and Gene Symbol. Institute of Genetic Resorces; Kyushu University: 2005.
-
- Cho S, Mitchell A, Regier JC, Mitter C, Poole RW, Friedlander TP, Zhao S. A highly conserved nuclear gene for low—level phylogenetics: elongation factor-1 alpha recovers morphology—based tree for heliothine moths. Molecular and Biological Evolution. 1995;12:650–656. - PubMed
-
- Daimon T, Hamada K, Mita K, Okano K, Suzuki MG, Kobayashi M, Shimada T. A Bombyx mori gene, BmChi-h, encodes a protein homologous to bacterial and baculovirus chitinases. Insect Biochemistry and Molecular Biology. 2003;33:749–759. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Miscellaneous

