Olive phenolic compounds: metabolic and transcriptional profiling during fruit development
- PMID: 22963618
- PMCID: PMC3480905
- DOI: 10.1186/1471-2229-12-162
Olive phenolic compounds: metabolic and transcriptional profiling during fruit development
Abstract
Background: Olive (Olea europaea L.) fruits contain numerous secondary metabolites, primarily phenolics, terpenes and sterols, some of which are particularly interesting for their nutraceutical properties. This study will attempt to provide further insight into the profile of olive phenolic compounds during fruit development and to identify the major genetic determinants of phenolic metabolism.
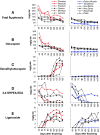
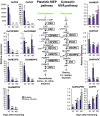
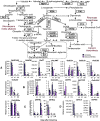
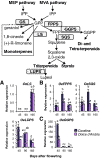
Results: The concentration of the major phenolic compounds, such as oleuropein, demethyloleuropein, 3-4 DHPEA-EDA, ligstroside, tyrosol, hydroxytyrosol, verbascoside and lignans, were measured in the developing fruits of 12 olive cultivars. The content of these compounds varied significantly among the cultivars and decreased during fruit development and maturation, with some compounds showing specificity for certain cultivars. Thirty-five olive transcripts homologous to genes involved in the pathways of the main secondary metabolites were identified from the massive sequencing data of the olive fruit transcriptome or from cDNA-AFLP analysis. Their mRNA levels were determined using RT-qPCR analysis on fruits of high- and low-phenolic varieties (Coratina and Dolce d'Andria, respectively) during three different fruit developmental stages. A strong correlation was observed between phenolic compound concentrations and transcripts putatively involved in their biosynthesis, suggesting a transcriptional regulation of the corresponding pathways. OeDXS, OeGES, OeGE10H and OeADH, encoding putative 1-deoxy-D-xylulose-5-P synthase, geraniol synthase, geraniol 10-hydroxylase and arogenate dehydrogenase, respectively, were almost exclusively present at 45 days after flowering (DAF), suggesting that these compounds might play a key role in regulating secoiridoid accumulation during fruit development.
Conclusions: Metabolic and transcriptional profiling led to the identification of some major players putatively involved in biosynthesis of secondary compounds in the olive tree. Our data represent the first step towards the functional characterisation of important genes for the determination of olive fruit quality.
Figures


References
-
- Oliveras López MJ, Innocenti M, Ieri F, Giaccherini C, Romani A, Mulinacci N. HPLC/DAD/ESI/MS detection of lignans from Spanish and Italian Olea europaea L. fruits. Journal of Food Composition and Analysis. 2008;21:62–70. doi: 10.1016/j.jfca.2007.04.012. - DOI
-
- Servili M, Selvaggini R, Esposto S, Taticchi A, Montedoro G, Morozzi G. Health and sensory properties of virgin olive oil hydrophilic phenols: agronomic and technological aspects of production that affect their occurrence in the oil. J Chromatogr A. 2004;1054(1–2):113–127. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

