Internal amino acids promote Gap1 permease ubiquitylation via TORC1/Npr1/14-3-3-dependent control of the Bul arrestin-like adaptors
- PMID: 22966204
- PMCID: PMC3486192
- DOI: 10.1128/MCB.00463-12
Internal amino acids promote Gap1 permease ubiquitylation via TORC1/Npr1/14-3-3-dependent control of the Bul arrestin-like adaptors
Abstract
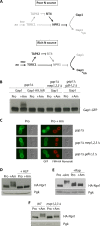
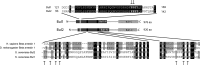
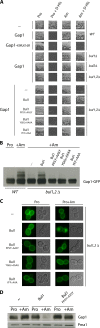
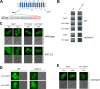
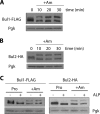
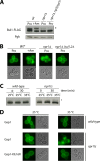
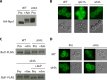
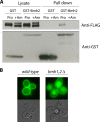
Ubiquitylation of many plasma membrane proteins promotes their endocytosis followed by degradation in the lysosome. The yeast general amino acid permease, Gap1, is ubiquitylated and downregulated when a good nitrogen source like ammonium is provided to cells growing on a poor nitrogen source. This ubiquitylation requires the Rsp5 ubiquitin ligase and the redundant arrestin-like Bul1 and Bul2 adaptors. Previous studies have shown that Gap1 ubiquitylation involves the TORC1 kinase complex, which inhibits the Sit4 phosphatase. This causes inactivation of the protein kinase Npr1, which protects Gap1 against ubiquitylation. However, the mechanisms inducing Gap1 ubiquitylation after Npr1 inactivation remain unknown. We here show that on a poor nitrogen source, the Bul adaptors are phosphorylated in an Npr1-dependent manner and bound to 14-3-3 proteins that protect Gap1 against downregulation. After ammonium is added and converted to amino acids, the Bul proteins are dephosphorylated, dissociate from the 14-3-3 proteins, and undergo ubiquitylation. Furthermore, dephosphorylation of Bul requires the Sit4 phosphatase, which is essential to Gap1 downregulation. The data support the emerging concept that permease ubiquitylation results from activation of the arrestin-like adaptors of the Rsp5 ubiquitin ligase, this coinciding with their dephosphorylation, dissociation from the inhibitory 14-3-3 proteins, and ubiquitylation.
Figures


Comment in
-
The running of the Buls: control of permease trafficking by α-arrestins Bul1 and Bul2.Mol Cell Biol. 2012 Nov;32(22):4506-9. doi: 10.1128/MCB.01176-12. Epub 2012 Oct 1. Mol Cell Biol. 2012. PMID: 23028041 Free PMC article. No abstract available.
Similar articles
-
Stress conditions promote yeast Gap1 permease ubiquitylation and down-regulation via the arrestin-like Bul and Aly proteins.J Biol Chem. 2014 Aug 8;289(32):22103-16. doi: 10.1074/jbc.M114.582320. Epub 2014 Jun 18. J Biol Chem. 2014. PMID: 24942738 Free PMC article.
-
Cooperative and selective roles of the WW domains of the yeast Nedd4-like ubiquitin ligase Rsp5 in the recognition of the arrestin-like adaptors Bul1 and Bul2.Biochem Biophys Res Commun. 2015 Jul 17-24;463(1-2):76-81. doi: 10.1016/j.bbrc.2015.05.025. Epub 2015 May 18. Biochem Biophys Res Commun. 2015. PMID: 25998383
-
The Bul1/2 Alpha-Arrestins Promote Ubiquitylation and Endocytosis of the Can1 Permease upon Cycloheximide-Induced TORC1-Hyperactivation.Int J Mol Sci. 2021 Sep 22;22(19):10208. doi: 10.3390/ijms221910208. Int J Mol Sci. 2021. PMID: 34638549 Free PMC article.
-
Ubiquitin ligase adaptors: regulators of ubiquitylation and endocytosis of plasma membrane proteins.Exp Cell Res. 2009 May 15;315(9):1574-83. doi: 10.1016/j.yexcr.2008.11.014. Epub 2008 Dec 3. Exp Cell Res. 2009. PMID: 19070615 Review.
-
The α-arrestin family of ubiquitin ligase adaptors links metabolism with selective endocytosis.Biol Cell. 2021 Apr;113(4):183-219. doi: 10.1111/boc.202000137. Epub 2021 Feb 1. Biol Cell. 2021. PMID: 33314196 Review.
Cited by
-
AMPK-Mediated Regulation of Alpha-Arrestins and Protein Trafficking.Int J Mol Sci. 2019 Jan 25;20(3):515. doi: 10.3390/ijms20030515. Int J Mol Sci. 2019. PMID: 30691068 Free PMC article. Review.
-
Aneuploidy shortens replicative lifespan in Saccharomyces cerevisiae.Aging Cell. 2016 Apr;15(2):317-24. doi: 10.1111/acel.12443. Epub 2016 Jan 13. Aging Cell. 2016. PMID: 26762766 Free PMC article.
-
A highly conserved mechanism for the detoxification and assimilation of the toxic phytoproduct L-azetidine-2-carboxylic acid in Aspergillus nidulans.Sci Rep. 2021 Apr 1;11(1):7391. doi: 10.1038/s41598-021-86622-3. Sci Rep. 2021. PMID: 33795709 Free PMC article.
-
FTY720/fingolimod, a sphingosine analogue, reduces amyloid-β production in neurons.PLoS One. 2013 May 7;8(5):e64050. doi: 10.1371/journal.pone.0064050. Print 2013. PLoS One. 2013. PMID: 23667698 Free PMC article.
-
Identification of the endocytic sorting signal recognized by the Art1-Rsp5 ubiquitin ligase complex.Mol Biol Cell. 2016 Dec 15;27(25):4043-4054. doi: 10.1091/mbc.E16-08-0570. Epub 2016 Oct 19. Mol Biol Cell. 2016. PMID: 27798240 Free PMC article.
References
-
- Béchet J, Grenson M, Wiame JM. 1970. Mutations affecting the repressibility of arginine biosynthetic enzymes in Saccharomyces cerevisiae. Eur. J. Biochem. 12:31–39 - PubMed
-
- Binda M, et al. 2009. The Vam6 GEF controls TORC1 by activating the EGO complex. Mol. Cell 35:563–573 - PubMed
-
- Boeckstaens M, Andre B, Marini AM. 2007. The yeast ammonium transport protein Mep2 and its positive regulator, the Npr1 kinase, play an important role in normal and pseudohyphal growth on various nitrogen media through retrieval of excreted ammonium. Mol. Microbiol. 64:534–546 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous
