Syntenic gene analysis between Brassica rapa and other Brassicaceae species
- PMID: 22969786
- PMCID: PMC3430884
- DOI: 10.3389/fpls.2012.00198
Syntenic gene analysis between Brassica rapa and other Brassicaceae species
Abstract
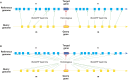
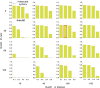
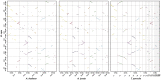
Chromosomal synteny analysis is important in genome comparison to reveal genomic evolution of related species. Shared synteny describes genomic fragments from different species that originated from an identical ancestor. Syntenic genes are orthologs located in these syntenic fragments, so they often share similar functions. Syntenic gene analysis is very important in Brassicaceae species to share gene annotations and investigate genome evolution. Here we designed and developed a direct and efficient tool, SynOrths, to identify pairwise syntenic genes between genomes of Brassicaceae species. SynOrths determines whether two genes are a conserved syntenic pair based not only on their sequence similarity, but also by the support of homologous flanking genes. Syntenic genes between Arabidopsis thaliana and Brassica rapa, Arabidopsis lyrata and B. rapa, and Thellungiella parvula and B. rapa were then identified using SynOrths. The occurrence of genome triplication in B. rapa was clearly observed, many genes that were evenly distributed in the genomes of A. thaliana, A. lyrata, and T. parvula had three syntenic copies in B. rapa. Additionally, there were many B. rapa genes that had no syntenic orthologs in A. thaliana, but some of these had syntenic orthologs in A. lyrata or T. parvula. Only 5,851 genes in B. rapa had no syntenic counterparts in any of the other three species. These 5,851 genes could have originated after B. rapa diverged from these species. A tool for syntenic gene analysis between species of Brassicaceae was developed, SynOrths, which could be used to accurately identify syntenic genes in differentiated but closely-related genomes. With this tool, we identified syntenic gene sets between B. rapa and each of A. thaliana, A. lyrata, T. parvula. Syntenic gene analysis is important for not only the gene annotation of newly sequenced Brassicaceae genomes by bridging them to model plant A. thaliana, but also the study of genome evolution in these species.
Keywords: Arabidopsis lyrata; Arabidopsis thaliana; Brassica rapa; Brassicaceae; Thellugiella parvula; ortholog; synteny.
Figures
References
-
- Hu T. T., Pattyn P., Bakker E. G., Cao J., Cheng J. F., Clark R. M., Fahlgren N., Fawcett J. A., Grimwood J., Gundlach H., Haberer G., Hollister J. D., Ossowski S., Ottilar R. P., Salamov A. A., Schneeberger K., Spannagl M., Wang X., Yang L., Nasrallah M. E., Bergelson J., Carrington J. C., Gaut B. S., Schmutz J., Mayer K. F., Van de Peer Y., Grigoriev I. V., Nordborg M., Weigel D., Guo Y. L. (2011). The Arabidopsis lyrata genome sequence and the basis of rapid genome size change. Nat. Genet. 43, 476–481 10.1038/ng.807 - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources

