A fluorescent, genetically-encoded voltage probe capable of resolving action potentials
- PMID: 22970127
- PMCID: PMC3435330
- DOI: 10.1371/journal.pone.0043454
A fluorescent, genetically-encoded voltage probe capable of resolving action potentials
Abstract
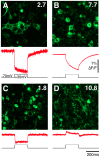
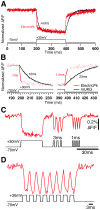
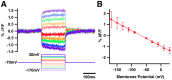
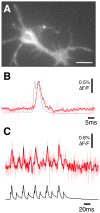
There is a pressing need in neuroscience for genetically-encoded, fluorescent voltage probes that can be targeted to specific neurons and circuits to allow study of neural activity using fluorescent imaging. We created 90 constructs in which the voltage sensing portion (S1-S4) of Ciona intestinalis voltage sensitive phosphatase (CiVSP) was fused to circularly permuted eGFP. This led to ElectricPk, a probe that is an order of magnitude faster (taus ~1-2 ms) than any currently published fluorescent protein-based voltage probe. ElectricPk can follow the rise and fall of neuronal action potentials with a modest decrease in fluorescence intensity (~0.7% ΔF/F). The probe has a nearly linear fluorescence/membrane potential response to both hyperpolarizing and depolarizing steps. This is the first probe based on CiVSP that captures the rapid movements of the voltage sensor, suggesting that voltage probes designed with circularly permuted fluorescent proteins may have some advantages.
Conflict of interest statement
Figures



References
-
- Siegel MS, Isacoff EY (1997) A genetically encoded optical probe of membrane voltage. Neuron 19: 735–741. - PubMed
-
- Sakai R, Repunte-Canonigo V, Raj CD, Knopfel T (2001) Design and characterization of a DNA-encoded, voltage-sensitive fluorescent protein. Eur J Neurosci 13: 2314–2318. - PubMed
-
- Tsutsui H, Karasawa S, Okamura Y, Miyawaki A (2008) Improving membrane voltage measurements using FRET with new fluorescent proteins. Nat Methods 5: 683–685. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

