Molecular modeling of the reaction pathway and hydride transfer reactions of HMG-CoA reductase
- PMID: 22971202
- PMCID: PMC3522576
- DOI: 10.1021/bi3008593
Molecular modeling of the reaction pathway and hydride transfer reactions of HMG-CoA reductase
Abstract
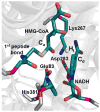
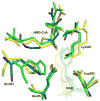
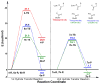
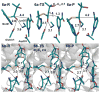
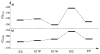
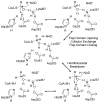
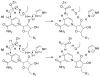
HMG-CoA reductase catalyzes the four-electron reduction of HMG-CoA to mevalonate and is an enzyme of considerable biomedical relevance because of the impact of its statin inhibitors on public health. Although the reaction has been studied extensively using X-ray crystallography, there are surprisingly no computational studies that test the mechanistic hypotheses suggested for this complex reaction. Theozyme and quantum mechanical (QM)/molecular mechanical (MM) calculations up to the B3LYP/6-31g(d,p)//B3LYP/6-311++g(2d,2p) level of theory were employed to generate an atomistic description of the enzymatic reaction process and its energy profile. The models generated here predict that the catalytically important Glu83 is protonated prior to hydride transfer and that it acts as the general acid or base in the reaction. With Glu83 protonated, the activation energies calculated for the sequential hydride transfer reactions, 21.8 and 19.3 kcal/mol, are in qualitative agreement with the experimentally determined rate constant for the entire reaction (1 s(-1) to 1 min(-1)). When Glu83 is not protonated, the first hydride transfer reaction is predicted to be disfavored by >20 kcal/mol, and the activation energy is predicted to be higher by >10 kcal/mol. While not involved in the reaction as an acid or base, Lys267 is critical for stabilization of the transition state in forming an oxyanion hole with the protonated Glu83. Molecular dynamics simulations and MM/Poisson-Boltzmann surface area free energy calculations predict that the enzyme active site stabilizes the hemithioacetal intermediate better than the aldehyde intermediate. This suggests a mechanism in which cofactor exchange occurs before the breakdown of the hemithioacetal. Slowing the conversion to aldehyde would provide the enzyme with a mechanism to protect it from solvent and explain why the free aldehyde is not observed experimentally. Our results support the hypothesis that the pK(a) of an active site acidic group is modulated by the redox state of the cofactor. The oxidized cofactor and deprotonated Glu83 are closer in space after hydride transfer, indicating that indeed the cofactor may influence the pK(a) of Glu83 through an electrostatic interaction. The enzyme is able to catalyze the transfer of a hydride to the structurally and electronically distinct substrates by maintaining the general shape of the active site and adjusting the electrostatic environment through acid-base chemistry. Our results are in good agreement with the well-studied hydride transfer reactions catalyzed by liver alcohol dehydrogenase in calculated energy profile and reaction geometries despite different mechanistic functionalities.
Figures

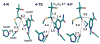

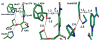

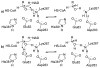
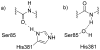
References
-
- Feng LL, Zhou L, Sun Y, Gui J, Wang XF, Wu P, Wan J, Ren YL, Qiu SX, Wei XY, Li J. Specific inhibitions of annonaceous acetogenins on class II 3-hydroxy-3-methylglutaryl coenzyme A reductase from Streptococcus pneumoniae. Bioorg Med Chem. 2011;19:3512–3519. - PubMed
-
- Li D, Gui J, Li Y, Feng L, Han X, Sun Y, Sun T, Chen Z, Cao Y, Zhang Y, Zhou L, Hu X, Ren Y, Wan J. Structure-Based Design and Screen of Novel Inhibitors for Class II 3-Hydroxy-3-methylglutaryl Coenzyme A Reductase from Streptococcus Pneumoniae. J Chem Inf Model. 2012;52:1833–1841. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

