Genetic ablation of arginase 1 in macrophages and neutrophils enhances clearance of an arthritogenic alphavirus
- PMID: 22972923
- PMCID: PMC3466331
- DOI: 10.4049/jimmunol.1201240
Genetic ablation of arginase 1 in macrophages and neutrophils enhances clearance of an arthritogenic alphavirus
Abstract
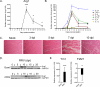
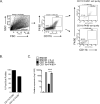
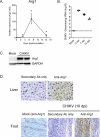
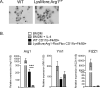
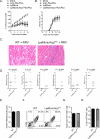
Chikungunya virus (CHIKV) and Ross River virus (RRV) cause a debilitating, and often chronic, musculoskeletal inflammatory disease in humans. Macrophages constitute the major inflammatory infiltrates in musculoskeletal tissues during these infections. However, the precise macrophage effector functions that affect the pathogenesis of arthritogenic alphaviruses have not been defined. We hypothesized that the severe damage to musculoskeletal tissues observed in RRV- or CHIKV-infected mice would promote a wound-healing response characterized by M2-like macrophages. Indeed, we found that RRV- and CHIKV-induced musculoskeletal inflammatory lesions, and macrophages present in these lesions, have a unique gene-expression pattern characterized by high expression of arginase 1 and Ym1/Chi3l3 in the absence of FIZZ1/Relmα that is consistent with an M2-like activation phenotype. Strikingly, mice specifically deleted for arginase 1 in neutrophils and macrophages had dramatically reduced viral loads and improved pathology in musculoskeletal tissues at late times post-RRV infection. These findings indicate that arthritogenic alphavirus infection drives a unique myeloid cell activation program in inflamed musculoskeletal tissues that inhibits virus clearance and impedes disease resolution in an arginase 1-dependent manner.
Figures




Similar articles
-
Myeloid Cell Arg1 Inhibits Control of Arthritogenic Alphavirus Infection by Suppressing Antiviral T Cells.PLoS Pathog. 2015 Oct 5;11(10):e1005191. doi: 10.1371/journal.ppat.1005191. eCollection 2015 Oct. PLoS Pathog. 2015. PMID: 26436766 Free PMC article.
-
Effects of an In-Frame Deletion of the 6k Gene Locus from the Genome of Ross River Virus.J Virol. 2016 Mar 28;90(8):4150-4159. doi: 10.1128/JVI.03192-15. Print 2016 Apr. J Virol. 2016. PMID: 26865723 Free PMC article.
-
Inflammatory monocytes mediate control of acute alphavirus infection in mice.PLoS Pathog. 2017 Dec 15;13(12):e1006748. doi: 10.1371/journal.ppat.1006748. eCollection 2017 Dec. PLoS Pathog. 2017. PMID: 29244871 Free PMC article.
-
Arthritogenic Alphavirus-Induced Immunopathology and Targeting Host Inflammation as A Therapeutic Strategy for Alphaviral Disease.Viruses. 2019 Mar 22;11(3):290. doi: 10.3390/v11030290. Viruses. 2019. PMID: 30909385 Free PMC article. Review.
-
Clinical and pathologic aspects of arthritis due to Ross River virus and other alphaviruses.Curr Opin Rheumatol. 2004 Jul;16(4):374-9. doi: 10.1097/01.bor.0000130537.76808.26. Curr Opin Rheumatol. 2004. PMID: 15201600 Review.
Cited by
-
Mechanisms underlying the pathogenesis of arthritogenic alphaviruses: host immune responses and virus persistence.Future Virol. 2019;9(5):441-444. doi: 10.2217/fvl.14.27. Epub 2014 Jun 16. Future Virol. 2019. PMID: 33907560 Free PMC article. No abstract available.
-
Neutrophil Functional Heterogeneity and Implications for Viral Infections and Treatments.Cells. 2022 Apr 13;11(8):1322. doi: 10.3390/cells11081322. Cells. 2022. PMID: 35456003 Free PMC article. Review.
-
C-type lectin receptor DCIR contributes to hippocampal injury in acute neurotropic virus infection.Sci Rep. 2021 Dec 10;11(1):23819. doi: 10.1038/s41598-021-03201-2. Sci Rep. 2021. PMID: 34893671 Free PMC article.
-
Disease Resolution in Chikungunya-What Decides the Outcome?Front Immunol. 2020 Apr 28;11:695. doi: 10.3389/fimmu.2020.00695. eCollection 2020. Front Immunol. 2020. PMID: 32411133 Free PMC article. Review.
-
Macrophage scavenger receptor 1 controls Chikungunya virus infection through autophagy in mice.Commun Biol. 2020 Oct 8;3(1):556. doi: 10.1038/s42003-020-01285-6. Commun Biol. 2020. PMID: 33033362 Free PMC article.
References
-
- Williams MC, Woodall JP, Gillett JD. O'nyong-Nyong Fever: An Epidemic Virus Diesease in East Africa. Vii. Virus Isolations from Man and Serological Studies up to July 1961. Trans R Soc Trop Med Hyg. 1965;59:186–197. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

