Stop codons in bacteria are not selectively equivalent
- PMID: 22974057
- PMCID: PMC3549826
- DOI: 10.1186/1745-6150-7-30
Stop codons in bacteria are not selectively equivalent
Abstract
Background: The evolution and genomic stop codon frequencies have not been rigorously studied with the exception of coding of non-canonical amino acids. Here we study the rate of evolution and frequency distribution of stop codons in bacterial genomes.
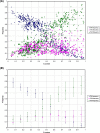
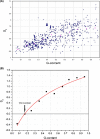
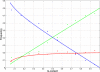
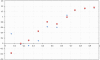
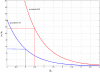
Results: We show that in bacteria stop codons evolve slower than synonymous sites, suggesting the action of weak negative selection. However, the frequency of stop codons relative to genomic nucleotide content indicated that this selection regime is not straightforward. The frequency of TAA and TGA stop codons is GC-content dependent, with TAA decreasing and TGA increasing with GC-content, while TAG frequency is independent of GC-content. Applying a formal, analytical model to these data we found that the relationship between stop codon frequencies and nucleotide content cannot be explained by mutational biases or selection on nucleotide content. However, with weak nucleotide content-dependent selection on TAG, -0.5 < Nes < 1.5, the model fits all of the data and recapitulates the relationship between TAG and nucleotide content. For biologically plausible rates of mutations we show that, in bacteria, TAG stop codon is universally associated with lower fitness, with TAA being the optimal for G-content < 16% while for G-content > 16% TGA has a higher fitness than TAG.
Conclusions: Our data indicate that TAG codon is universally suboptimal in the bacterial lineage, such that TAA is likely to be the preferred stop codon for low GC content while the TGA is the preferred stop codon for high GC content. The optimization of stop codon usage may therefore be useful in genome engineering or gene expression optimization applications.
Figures
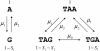

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

