Annotation of the modular polyketide synthase and nonribosomal peptide synthetase gene clusters in the genome of Streptomyces tsukubaensis NRRL18488
- PMID: 22983969
- PMCID: PMC3497360
- DOI: 10.1128/AEM.01891-12
Annotation of the modular polyketide synthase and nonribosomal peptide synthetase gene clusters in the genome of Streptomyces tsukubaensis NRRL18488
Abstract
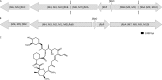
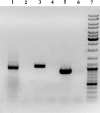
The high G+C content and large genome size make the sequencing and assembly of Streptomyces genomes more difficult than for other bacteria. Many pharmaceutically important natural products are synthesized by modular polyketide synthases (PKSs) and nonribosomal peptide synthetases (NRPSs). The analysis of such gene clusters is difficult if the genome sequence is not of the highest quality, because clusters can be distributed over several contigs, and sequencing errors can introduce apparent frameshifts into the large PKS and NRPS proteins. An additional problem is that the modular nature of the clusters results in the presence of imperfect repeats, which may cause assembly errors. The genome sequence of Streptomyces tsukubaensis NRRL18488 was scanned for potential PKS and NRPS modular clusters. A phylogenetic approach was used to identify multiple contigs belonging to the same cluster. Four PKS clusters and six NRPS clusters were identified. Contigs containing cluster sequences were analyzed in detail by using the ClustScan program, which suggested the order and orientation of the contigs. The sequencing of the appropriate PCR products confirmed the ordering and allowed the correction of apparent frameshifts resulting from sequencing errors. The product chemistry of such correctly assembled clusters could also be predicted. The analysis of one PKS cluster showed that it should produce a bafilomycin-like compound, and reverse transcription (RT)-PCR was used to show that the cluster was transcribed.
Figures


References
-
- Baltz RH. 2006. Marcel Faber Roundtable: is our antibiotic pipeline unproductive because of starvation, constipation or lack of inspiration? J. Ind. Microbiol. Biotechnol. 33:507–513 - PubMed
-
- Bentley SD, et al. 2002. Complete genome sequence of the model actinomycete Streptomyces coelicolor A3(2). Nature 417:141–147 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Miscellaneous

