Sf-PHB2, a new transcription factor, drives WSSV Ie1 gene expression via a 12-bp DNA element
- PMID: 22985503
- PMCID: PMC3511282
- DOI: 10.1186/1743-422X-9-206
Sf-PHB2, a new transcription factor, drives WSSV Ie1 gene expression via a 12-bp DNA element
Abstract
Background: The WSSV immediate early gene ie1 is highly expressed throughout viral infection cycle and may play a central role in initiating viral replication during infection.
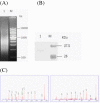
Results: Here, a detailed characterization of the ie1 promoter was performed using deletion and mutation analyses to elucidate the role of the individual promoter motifs. Three results were obtained: 1) the ie1 promoter is a classical eukaryotic promoter that contains the initiator element (Inr) and TATA box responsible for the basal promoter activity; 2) mutation or truncation of a predicted Sp1 site decreased the level of promoter activity by about 3-fold, indicating that the Sp1 site is an important cis-element of the promoter; and 3) truncation of a 12-bp sequence that resides at -78/-67 of the ie1 promoter decreased the level of promoter activity by about 14-fold, indicating that the 12-bp motif is a critical upstream element of the ie1 promoter for binding of a strong transcription factor to drive the ie1 gene expression in the cells. Further, the 12-bp DNA binding protein was purified from the nuclear proteins of Sf9 cells using DNA affinity chromatography, and was identified as a homologue of the prohibitin2 protein (named as Sf-PHB2) using mass spectrometry. Furthermore, the DNA binding activity of Sf-PHB2 was verified using a super shift analysis.
Conclusion: These results support that the Sf-PHB2 is a novel transcription factor that drives WSSV ie1 gene expression by binding to the 12-bp DNA element.
Figures




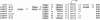
References
-
- Chou HY, Huang CY, Wang CH, Chiang HC, Lo CF. Pathogenicity of a baculovirus infection causing white spot syndrome in cultured penaeid shrimp in Taiwan. Dis Aquat Org. 1995;23:165–173.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

