Understanding the origins of UV-induced recombination through manipulation of sister chromatid cohesion
- PMID: 22987150
- PMCID: PMC3507489
- DOI: 10.4161/cc.21945
Understanding the origins of UV-induced recombination through manipulation of sister chromatid cohesion
Abstract
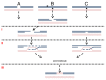
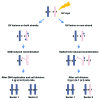
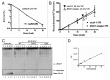
Ultraviolet light (UV) can provoke genome instability, partly through its ability to induce homologous recombination (HR). However, the mechanism(s) of UV-induced recombination is poorly understood. Although double-strand breaks (DSBs) have been invoked, there is little evidence for their generation by UV. Alternatively, single-strand DNA lesions that stall replication forks could provoke recombination. Recent findings suggest efficient initiation of UV-induced recombination in G1 through processing of closely spaced single-strand lesions to DSBs. However, other scenarios are possible, since the recombination initiated in G1 can be completed in the following stages of the cell cycle. We developed a system that could address UV-induced recombination events that start and finish in G2 by manipulating the activity of the sister chromatid cohesion complex. Here we show that sister-chromatid cohesion suppresses UV-induced recombination events that are initiated and resolved in G2. By comparing recombination frequencies and survival between UV and ionizing radiation, we conclude that a substantial portion of UV-induced recombination occurs through DSBs. This notion is supported by a direct physical observation of UV-induced DSBs that are dependent on nucleotide excision repair. However, a significant role of nonDSB intermediates in UV-induced recombination cannot be excluded.
Figures

References
-
- Andersen SL, Sekelsky J. Meiotic versus mitotic recombination: two different routes for double-strand break repair: the different functions of meiotic versus mitotic DSB repair are reflected in different pathway usage and different outcomes. Bioessays. 2010;32:1058–66. doi: 10.1002/bies.201000087. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
