Binding of two DNA molecules by type II topoisomerases for decatenation
- PMID: 22989710
- PMCID: PMC3510509
- DOI: 10.1093/nar/gks843
Binding of two DNA molecules by type II topoisomerases for decatenation
Abstract
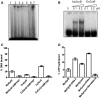
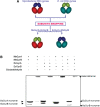
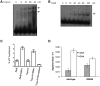
Topoisomerases (topos) maintain DNA topology and influence DNA transaction processes by catalysing relaxation, supercoiling and decatenation reactions. In the cellular milieu, division of labour between different topos ensures topological homeostasis and control of central processes. In Escherichia coli, DNA gyrase is the principal enzyme that carries out negative supercoiling, while topo IV catalyses decatenation, relaxation and unknotting. DNA gyrase apparently has the daunting task of undertaking both the enzyme functions in mycobacteria, where topo IV is absent. We have shown previously that mycobacterial DNA gyrase is an efficient decatenase. Here, we demonstrate that the strong decatenation property of the enzyme is due to its ability to capture two DNA segments in trans. Topo IV, a strong dedicated decatenase of E. coli, also captures two distinct DNA molecules in a similar manner. In contrast, E. coli DNA gyrase, which is a poor decatenase, does not appear to be able to hold two different DNA molecules in a stable complex. The binding of a second DNA molecule to GyrB/ParE is inhibited by ATP and the non-hydrolysable analogue, AMPPNP, and by the substitution of a prominent positively charged residue in the GyrB N-terminal cavity, suggesting that this binding represents a potential T-segment positioned in the cavity. Thus, after the GyrA/ParC mediated initial DNA capture, GyrB/ParE would bind efficiently to a second DNA in trans to form a T-segment prior to nucleotide binding and closure of the gate during decatenation.
Figures




References
-
- Filutowicz M. Requirement of DNA gyrase for the initiation of chromosome replication in Escherichia coli K-12. Mol. Gen. Genet. 1980;177:301–309. - PubMed
-
- Masse E, Drolet M. Relaxation of transcription-induced negative supercoiling is an essential function of Escherichia coli DNA topoisomerase I. J. Biol. Chem. 1999;274:16654–16658. - PubMed
-
- Zechiedrich EL, Cozzarelli NR. Roles of topoisomerase IV and DNA gyrase in DNA unlinking during replication in Escherichia coli. Genes Dev. 1995;9:2859–2869. - PubMed

