G protein-coupled receptor-type G proteins are required for light-dependent seedling growth and fertility in Arabidopsis
- PMID: 23001037
- PMCID: PMC3480293
- DOI: 10.1105/tpc.112.098681
G protein-coupled receptor-type G proteins are required for light-dependent seedling growth and fertility in Arabidopsis
Abstract
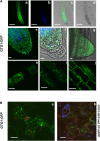
G protein-coupled receptor-type G proteins (GTGs) are highly conserved membrane proteins in plants, animals, and fungi that have eight to nine predicted transmembrane domains. They have been classified as G protein-coupled receptor-type G proteins that function as abscisic acid (ABA) receptors in Arabidopsis thaliana. We cloned Arabidopsis GTG1 and GTG2 and isolated new T-DNA insertion alleles of GTG1 and GTG2 in both Wassilewskija and Columbia backgrounds. These gtg1 gtg2 double mutants show defects in fertility, hypocotyl and root growth, and responses to light and sugars. Histological studies of shoot tissue reveal cellular distortions that are particularly evident in the epidermal layer. Stable expression of GTG1(pro):GTG1-GFP (for green fluorescent protein) in Arabidopsis and transient expression in tobacco (Nicotiana tabacum) indicate that GTG1 is localized primarily to Golgi bodies and to the endoplasmic reticulum. Microarray analysis comparing gene expression profiles in the wild type and double mutant revealed differences in expression of genes important for cell wall function, hormone response, and amino acid metabolism. The double mutants isolated here respond normally to ABA in seed germination assays, root growth inhibition, and gene expression analysis. These results are inconsistent with their proposed role as ABA receptors but demonstrate that GTGs are fundamentally important for plant growth and development.
Figures











Similar articles
-
Genetic characterization reveals no role for the reported ABA receptor, GCR2, in ABA control of seed germination and early seedling development in Arabidopsis.Plant J. 2007 Dec;52(6):1001-13. doi: 10.1111/j.1365-313X.2007.03291.x. Epub 2007 Sep 25. Plant J. 2007. PMID: 17894782
-
Two novel GPCR-type G proteins are abscisic acid receptors in Arabidopsis.Cell. 2009 Jan 9;136(1):136-48. doi: 10.1016/j.cell.2008.12.026. Cell. 2009. PMID: 19135895
-
Arabidopsis SAG protein containing the MDN1 domain participates in seed germination and seedling development by negatively regulating ABI3 and ABI5.J Exp Bot. 2014 Jan;65(1):35-45. doi: 10.1093/jxb/ert343. Epub 2013 Oct 25. J Exp Bot. 2014. PMID: 24163287 Free PMC article.
-
From the Outside to the Inside: New Insights on the Main Factors That Guide Seed Dormancy and Germination.Genes (Basel). 2020 Dec 31;12(1):52. doi: 10.3390/genes12010052. Genes (Basel). 2020. PMID: 33396410 Free PMC article. Review.
-
Abscisic acid signaling in seeds and seedlings.Plant Cell. 2002;14 Suppl(Suppl):S15-45. doi: 10.1105/tpc.010441. Plant Cell. 2002. PMID: 12045268 Free PMC article. Review. No abstract available.
Cited by
-
Characterization of ZmCOLD1, novel GPCR-Type G Protein genes involved in cold stress from Zea mays L. and the evolution analysis with those from other species.Physiol Mol Biol Plants. 2021 Mar;27(3):619-632. doi: 10.1007/s12298-021-00966-8. Epub 2021 Mar 15. Physiol Mol Biol Plants. 2021. PMID: 33854288 Free PMC article.
-
Arabidopsis heterotrimeric G protein β subunit, AGB1, regulates brassinosteroid signalling independently of BZR1.J Exp Bot. 2013 Aug;64(11):3213-23. doi: 10.1093/jxb/ert159. Epub 2013 Jun 27. J Exp Bot. 2013. PMID: 23814276 Free PMC article.
-
A Sugarcane G-Protein-Coupled Receptor, ShGPCR1, Confers Tolerance to Multiple Abiotic Stresses.Front Plant Sci. 2021 Nov 11;12:745891. doi: 10.3389/fpls.2021.745891. eCollection 2021. Front Plant Sci. 2021. PMID: 35295863 Free PMC article.
-
Heterotrimeric G Protein-Regulated Ca2+ Influx and PIN2 Asymmetric Distribution Are Involved in Arabidopsis thaliana Roots' Avoidance Response to Extracellular ATP.Front Plant Sci. 2017 Sep 1;8:1522. doi: 10.3389/fpls.2017.01522. eCollection 2017. Front Plant Sci. 2017. PMID: 28919907 Free PMC article.
-
The key clock component ZEITLUPE (ZTL) negatively regulates ABA signaling by degradation of CHLH in Arabidopsis.Front Plant Sci. 2022 Sep 13;13:995907. doi: 10.3389/fpls.2022.995907. eCollection 2022. Front Plant Sci. 2022. PMID: 36176682 Free PMC article.
References
-
- Abe M., Katsumata H., Komeda Y., Takahashi T. (2003). Regulation of shoot epidermal cell differentiation by a pair of homeodomain proteins in Arabidopsis. Development 130: 635–643 - PubMed
-
- Barrero J.M., Piqueras P., González-Guzmán M., Serrano R., Rodríguez P.L., Ponce M.R., Micol J.L. (2005). A mutational analysis of the ABA1 gene of Arabidopsis thaliana highlights the involvement of ABA in vegetative development. J. Exp. Bot. 56: 2071–2083 - PubMed
-
- Boavida L.C., McCormick S. (2007). Temperature as a determinant factor for increased and reproducible in vitro pollen germination in Arabidopsis thaliana. Plant J. 52: 570–582 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

