Surviving in a toxic world: transcriptomics and gene expression profiling in response to environmental pollution in the critically endangered European eel
- PMID: 23009661
- PMCID: PMC3532374
- DOI: 10.1186/1471-2164-13-507
Surviving in a toxic world: transcriptomics and gene expression profiling in response to environmental pollution in the critically endangered European eel
Abstract
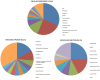
Background: Genomic and transcriptomic approaches have the potential for unveiling the genome-wide response to environmental perturbations. The abundance of the catadromous European eel (Anguilla anguilla) stock has been declining since the 1980s probably due to a combination of anthropogenic and climatic factors. In this paper, we explore the transcriptomic dynamics between individuals from high (river Tiber, Italy) and low pollution (lake Bolsena, Italy) environments, which were measured for 36 PCBs, several organochlorine pesticides and brominated flame retardants and nine metals.
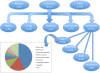
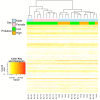
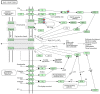
Results: To this end, we first (i) updated the European eel transcriptome using deep sequencing data with a total of 640,040 reads assembled into 44,896 contigs (Eeelbase release 2.0), and (ii) developed a transcriptomic platform for global gene expression profiling in the critically endangered European eel of about 15,000 annotated contigs, which was applied to detect differentially expressed genes between polluted sites. Several detoxification genes related to metabolism of pollutants were upregulated in the highly polluted site, including genes that take part in phase I of the xenobiotic metabolism (CYP3A), phase II (glutathione-S-transferase) and oxidative stress (glutathione peroxidase). In addition, key genes in the mitochondrial respiratory chain and oxidative phosphorylation were down-regulated at the Tiber site relative to the Bolsena site.
Conclusions: Together with the induced high expression of detoxification genes, the suggested lowered expression of genes supposedly involved in metabolism suggests that pollution may also be associated with decreased respiratory and energy production.
Figures



Similar articles
-
Detecting genome-wide gene transcription profiles associated with high pollution burden in the critically endangered European eel.Aquat Toxicol. 2013 May 15;132-133:157-64. doi: 10.1016/j.aquatox.2013.02.012. Epub 2013 Mar 4. Aquat Toxicol. 2013. PMID: 23518471
-
Gene transcription reflects poor health status of resident European eel chronically exposed to environmental pollutants.Aquat Toxicol. 2013 Jan 15;126:242-55. doi: 10.1016/j.aquatox.2012.11.006. Epub 2012 Nov 21. Aquat Toxicol. 2013. PMID: 23247545
-
Sequencing, de novo annotation and analysis of the first Anguilla anguilla transcriptome: EeelBase opens new perspectives for the study of the critically endangered European eel.BMC Genomics. 2010 Nov 16;11:635. doi: 10.1186/1471-2164-11-635. BMC Genomics. 2010. PMID: 21080939 Free PMC article.
-
The impact of chemical pollution on the European eel (Anguilla anguilla) from a Mediterranean hypersaline coastal lagoon.Environ Sci Pollut Res Int. 2023 Jul;30(33):80106-80122. doi: 10.1007/s11356-023-27871-9. Epub 2023 Jun 8. Environ Sci Pollut Res Int. 2023. PMID: 37289386 Free PMC article.
-
Gonadal transcriptome analysis of wild contaminated female European eels during artificial gonad maturation.Chemosphere. 2015 Nov;139:303-9. doi: 10.1016/j.chemosphere.2015.06.007. Epub 2015 Jul 6. Chemosphere. 2015. PMID: 26159298
Cited by
-
Transcriptome profile analysis reveals specific signatures of pollutants in Atlantic eels.Ecotoxicology. 2015 Jan;24(1):71-84. doi: 10.1007/s10646-014-1356-x. Epub 2014 Sep 26. Ecotoxicology. 2015. PMID: 25258179
-
In absence of local adaptation, plasticity and spatially varying selection rule: a view from genomic reaction norms in a panmictic species (Anguilla rostrata).BMC Genomics. 2014 May 27;15:403. doi: 10.1186/1471-2164-15-403. BMC Genomics. 2014. PMID: 24884429 Free PMC article.
-
Exposure to environmental radionuclides is associated with altered metabolic and immunity pathways in a wild rodent.Mol Ecol. 2019 Oct;28(20):4620-4635. doi: 10.1111/mec.15241. Epub 2019 Sep 30. Mol Ecol. 2019. PMID: 31498518 Free PMC article.
-
Transcriptome sequencing based annotation and homologous evidence based scaffolding of Anguilla japonica draft genome.BMC Genomics. 2016 Jan 11;17 Suppl 1(Suppl 1):13. doi: 10.1186/s12864-015-2306-6. BMC Genomics. 2016. PMID: 26818233 Free PMC article.
-
Detecting the exposure to Cd and PCBs by means of a non-invasive transcriptomic approach in laboratory and wild contaminated European eels (Anguilla anguilla).Environ Sci Pollut Res Int. 2016 Mar;23(6):5431-41. doi: 10.1007/s11356-015-5754-2. Epub 2015 Nov 14. Environ Sci Pollut Res Int. 2016. PMID: 26566612
References
-
- Travers SE, Smith MD, Bai J, Hulbert SH, Leach JE, Schnable PS, Knapp AK, Milliken GA, Fray PA, Saleh A, Garrett KA. Ecological genomics: making the leap from model systems in the lab to native populations in the field. Front Ecol Environ. 2007;5:19–24. doi: 10.1890/1540-9295(2007)5[19:EGMTLF]2.0.CO;2. - DOI
-
- Vandenkoornhuyse P, Dufresne A, Quaiser A, Gouesbet G, Binet F, Francez AJ, Mahé S, Bormans M, Lagadeux Y, Couée I. Integration of molecular functions at the ecosystemic level: breakthroughs and future goals of environmental genomics and post-genomics. Ecol Lett. 2010;13:776–791. doi: 10.1111/j.1461-0248.2010.01464.x. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

