Conditional disorder in chaperone action
- PMID: 23018052
- PMCID: PMC3508372
- DOI: 10.1016/j.tibs.2012.08.006
Conditional disorder in chaperone action
Abstract
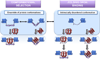
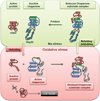
Protein disorder remains an intrinsically fuzzy concept. Its role in protein function is difficult to conceptualize and its experimental study is challenging. Although a wide variety of roles for protein disorder have been proposed, establishing that disorder is functionally important, particularly in vivo, is not a trivial task. Several molecular chaperones have now been identified as conditionally disordered proteins; fully folded and chaperone-inactive under non-stress conditions, they adopt a partially disordered conformation upon exposure to distinct stress conditions. This disorder appears to be vital for their ability to bind multiple aggregation-sensitive client proteins and to protect cells against the stressors. The study of these conditionally disordered chaperones should prove useful in understanding the functional role for protein disorder in molecular recognition.
Copyright © 2012 Elsevier Ltd. All rights reserved.
Figures
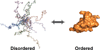

References
-
- Le Gall T, et al. Intrinsic disorder in the Protein Data Bank. Journal of biomolecular structure & dynamics. 2007;24:325–342. - PubMed
-
- Uversky VN. Intrinsically disordered proteins from A to Z. Int J Biochem Cell Biol. 2011;43:1090–1103. - PubMed
-
- Tompa P, Fuxreiter M. Fuzzy complexes: polymorphism and structural disorder in protein-protein interactions. Trends Biochem Sci. 2008;33:2–8. - PubMed
-
- Tompa P, et al. Prevalent structural disorder in E. coli and S. cerevisiae proteomes. J Proteome Res. 2006;5:1996–2000. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

