Non-cell autonomous control of apoptosis by ligand-independent Hedgehog signaling in Drosophila
- PMID: 23018595
- PMCID: PMC3554335
- DOI: 10.1038/cdd.2012.126
Non-cell autonomous control of apoptosis by ligand-independent Hedgehog signaling in Drosophila
Abstract
Hedgehog (Hh) signaling is important for development and homeostasis in vertebrates and invertebrates. Ligand-independent, deregulated Hh signaling caused by loss of negative regulators such as Patched causes excessive cell proliferation, leading to overgrowth in Drosophila and tumors in humans, including basal-cell carcinoma and medulloblastoma. We show that in Drosophila deregulated Hh signaling also promotes cell survival by increasing the resistance to apoptosis. Surprisingly, cells with deregulated Hh activity do not protect themselves from apoptosis; instead, they promote cell survival of neighboring wild-type cells. This non-cell autonomous effect is mediated by Hh-induced Notch signaling, which elevates the protein levels of Drosophila inhibitor of apoptosis protein-1 (Diap-1), conferring resistance to apoptosis. In summary, we demonstrate that deregulated Hh signaling not only promotes proliferation but also cell survival of neighboring cells. This non-cell autonomous control of apoptosis highlights an underappreciated function of deregulated Hh signaling, which may help to generate a supportive micro-environment for tumor development.
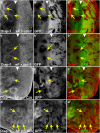
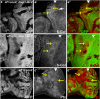
Figures




Similar articles
-
Ligand-independent activation of the Hedgehog pathway displays non-cell autonomous proliferation during eye development in Drosophila.Mech Dev. 2012 Jul;129(5-8):98-108. doi: 10.1016/j.mod.2012.05.009. Epub 2012 Jun 5. Mech Dev. 2012. PMID: 22677792 Free PMC article.
-
Notch signaling activates Yorkie non-cell autonomously in Drosophila.PLoS One. 2012;7(6):e37615. doi: 10.1371/journal.pone.0037615. Epub 2012 Jun 5. PLoS One. 2012. PMID: 22679484 Free PMC article.
-
Hedgehog-stimulated stem cells depend on non-canonical activity of the Notch co-activator Mastermind.Development. 2009 Jul;136(13):2177-86. doi: 10.1242/dev.035329. Epub 2009 May 27. Development. 2009. PMID: 19474148 Free PMC article.
-
Regulation of Hedgehog signaling: a complex story.Biochem Pharmacol. 2004 Mar 1;67(5):805-14. doi: 10.1016/j.bcp.2004.01.002. Biochem Pharmacol. 2004. PMID: 15104233 Free PMC article. Review.
-
Transducing the Hedgehog signal across the plasma membrane.Fly (Austin). 2007 Nov-Dec;1(6):333-6. doi: 10.4161/fly.5570. Epub 2007 Nov 15. Fly (Austin). 2007. PMID: 18820483 Review.
Cited by
-
Non-autonomous consequences of cell death and other perks of being metazoan.AIMS Genet. 2015;2(1):54-69. doi: 10.3934/genet.2015.1.54#sthash.dNy9tFhS.dpuf. AIMS Genet. 2015. PMID: 26069889 Free PMC article.
-
Neural stem cell progeny regulate stem cell death in a Notch and Hox dependent manner.Cell Death Differ. 2015 Aug;22(8):1378-87. doi: 10.1038/cdd.2014.235. Epub 2015 Jan 30. Cell Death Differ. 2015. PMID: 25633198 Free PMC article.
-
Hedgehog is a positive regulator of FGF signalling during embryonic tracheal cell migration.PLoS One. 2014 Mar 20;9(3):e92682. doi: 10.1371/journal.pone.0092682. eCollection 2014. PLoS One. 2014. PMID: 24651658 Free PMC article.
-
Multiple mechanisms modulate distinct cellular susceptibilities toward apoptosis in the developing Drosophila eye.Dev Cell. 2014 Jul 14;30(1):48-60. doi: 10.1016/j.devcel.2014.05.007. Epub 2014 Jun 26. Dev Cell. 2014. PMID: 24981611 Free PMC article.
-
The histone demethylase Kdm5 controls Hid-induced cell death in Drosophila.Front Cell Death. 2024;3:1471050. doi: 10.3389/fceld.2024.1471050. Epub 2024 Nov 19. Front Cell Death. 2024. PMID: 40416947
References
-
- Kumar S. Caspase function in programmed cell death. Cell Death Differ. 2007;14:32–43. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

