Application of a rapid, simple, and accurate adenovirus-based method to compare PET reporter gene/PET reporter probe systems
- PMID: 23054556
- PMCID: PMC3833443
- DOI: 10.1007/s11307-012-0596-5
Application of a rapid, simple, and accurate adenovirus-based method to compare PET reporter gene/PET reporter probe systems
Abstract
Purpose: This study aims to use a simple, quantitative method to compare the HSV1sr39TK/(18) F-FHBG PET reporter gene/PET reporter probe (PRG/PRP) system with PRGs derived from human nucleoside kinases.
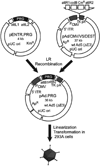
Procedures: The same adenovirus vector is used to express alternative PRGs. Equal numbers of vectors are injected intravenously into mice. After PRP imaging, quantitative hepatic PET signals are normalized for transduction by measuring hepatic viral genomes.
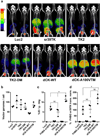
Results: The same adenovirus vector was used to express equivalent amounts of HSV1sr39TK, mutant human thymidine kinase 2 (TK2-DM), and mutant human deoxycytidine kinase (dCK-A100VTM) in mouse liver. HSV1sr39TK expression was measured with (18) F-FHBG, TK2-DM and dCK-A100VTM with (18) F-L-FMAU. TK2-DM/(18) F-L-FMAU and HSV1sr39TK/(18) F-FHBG had equivalent sensitivities; dCK-A100VTM/(18) F-L-FMAU was twice as sensitive as HSV1sr39TK/(18) F-FHBG.
Conclusions: The human PRG/PRP sensitivities are comparable and/or better than HSV1sr39TK/(18) F-FHBG. However, for clinical use, identification of the best PRP substrate for each enzyme, characterization of probe distribution, and consequences of overexpressing nucleoside kinases must be evaluated.
Conflict of interest statement
The authors declare they have no conflict of interest.
Figures

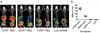
References
-
- Bhaumik S, Gambhir SS. Optical imaging of Renilla luciferase, synthetic Renilla luciferase, and firefly luciferase reporter gene expression in living mice. Journal of Biomedical Optics. 2004;9:578–586. - PubMed
-
- Gheysens O, Gambhir SS. Studying molecular and cellular processes in the intact organism. Progress in Drug Research. 2005;62:117–150. - PubMed
-
- Lewis Js, Achilefu S, Garbow JR, Laforest R, Welch MJ. Small animal imaging. Current technology and perspectives for oncological imaging. European Journal of Cancer. 2002;38:2173–2188. - PubMed
-
- Jenkins DE, Oei Y, Hornig YS, Yu S-F, Dusich J, Purchio T, Contag PR. Bioluminescent imaging (BLI) to improve and refine traditional murine models of tumor growth and metastasis. Clinical & Experimental Metastasis. 2003;20:733–744. - PubMed
-
- Iyer M, Sato M, Johnson M, Gambhir SS, Wu L. Applications of molecular imaging in cancer gene therapy. Current Gene Therapy. 2005;5:607–618. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

