High density GWAS for LDL cholesterol in African Americans using electronic medical records reveals a strong protective variant in APOE
- PMID: 23067351
- PMCID: PMC3521536
- DOI: 10.1111/j.1752-8062.2012.00446.x
High density GWAS for LDL cholesterol in African Americans using electronic medical records reveals a strong protective variant in APOE
Abstract
Only one low-density lipoprotein cholesterol (LDL-C) genome-wide association study (GWAS) has been previously reported in -African Americans. We performed a GWAS of LDL-C in African Americans using data extracted from electronic medical records (EMR) in the eMERGE network. African Americans were genotyped on the Illumina 1M chip. All LDL-C measurements, prescriptions, and diagnoses of concomitant disease were extracted from EMR. We created two analytic datasets; one dataset having median LDL-C calculated after the exclusion of some lab values based on comorbidities and medication (n= 618) and another dataset having median LDL-C calculated without any exclusions (n= 1,249). SNP rs7412 in APOE was strongly associated with LDL-C in both datasets (p < 5 × 10(-8) ). In the dataset with exclusions, a decrease of 20.0 mg/dL per minor allele was observed. The effect size was attenuated (12.3 mg/dL) in the dataset without any lab values excluded. Although other signals in APOE have been detected in previous GWAS, this large and important SNP association has not been well detected in large GWAS because rs7412 was not included on many genotyping arrays. Use of median LDL-C extracted from EMR after exclusions for medications and comorbidities increased the percentage of trait variance explained by genetic variation.
© 2012 Wiley Periodicals, Inc.
Figures

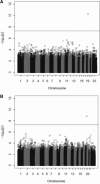
References
-
- Hindorff LA, Junkins HA, hall PN, Mehta JP, Manolio TA. A Catalog of Published Genome‐Wide Association Studies. 2010. Available at: http://www.genome.gov/gwastudies. Accessed on August 24, 2010.
Publication types
MeSH terms
Substances
Grants and funding
- U01-HG-004610/HG/NHGRI NIH HHS/United States
- UL1 TR000150/TR/NCATS NIH HHS/United States
- U01 HG004603/HG/NHGRI NIH HHS/United States
- U01HG004609/HG/NHGRI NIH HHS/United States
- U01 HG004609/HG/NHGRI NIH HHS/United States
- U01 HG004599/HG/NHGRI NIH HHS/United States
- U01 HG006378/HG/NHGRI NIH HHS/United States
- U01 HG004610/HG/NHGRI NIH HHS/United States
- U01-HG-004608/HG/NHGRI NIH HHS/United States
- U01 HG004608/HG/NHGRI NIH HHS/United States
- U01-HG-04599/HG/NHGRI NIH HHS/United States
- U01 HG006375/HG/NHGRI NIH HHS/United States
- KL2 RR025740/RR/NCRR NIH HHS/United States
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

