Distinguishing between selective sweeps from standing variation and from a de novo mutation
- PMID: 23071458
- PMCID: PMC3469416
- DOI: 10.1371/journal.pgen.1003011
Distinguishing between selective sweeps from standing variation and from a de novo mutation
Abstract
An outstanding question in human genetics has been the degree to which adaptation occurs from standing genetic variation or from de novo mutations. Here, we combine several common statistics used to detect selection in an Approximate Bayesian Computation (ABC) framework, with the goal of discriminating between models of selection and providing estimates of the age of selected alleles and the selection coefficients acting on them. We use simulations to assess the power and accuracy of our method and apply it to seven of the strongest sweeps currently known in humans. We identify two genes, ASPM and PSCA, that are most likely affected by selection on standing variation; and we find three genes, ADH1B, LCT, and EDAR, in which the adaptive alleles seem to have swept from a new mutation. We also confirm evidence of selection for one further gene, TRPV6. In one gene, G6PD, neither neutral models nor models of selective sweeps fit the data, presumably because this locus has been subject to balancing selection.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
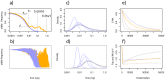
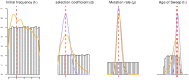


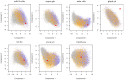
References
-
- Kimura M (1985) The Neutral Theory of Molecular Evolution. Cambridge University Press 388.
-
- Ohta T (1992) The nearly neutral theory of molecular evolution. Annual Review of Ecology and Systematics 23: 263–286.
-
- Hurst LD (2009) Genetics and the understanding of selection. Nat Rev Genet 10: 83–93 doi:10.1038/nrg2506 - DOI - PubMed
-
- Hernandez RD, Kelley JL, Elyashiv E, Melton SC, Auton A, et al. (2011) Classic Selective Sweeps Were Rare in Recent Human Evolution. Science 331: 920–924 doi:10.1126/science.1198878 - DOI - PMC - PubMed
-
- Hermisson J, Pennings PS (2005) Soft Sweeps. Genetics 169: 2335–2352 doi:10.1534/genetics.104.036947 - DOI - PMC - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

