Mechanistic study of broadly neutralizing human monoclonal antibodies against dengue virus that target the fusion loop
- PMID: 23077306
- PMCID: PMC3536401
- DOI: 10.1128/JVI.02273-12
Mechanistic study of broadly neutralizing human monoclonal antibodies against dengue virus that target the fusion loop
Abstract
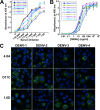
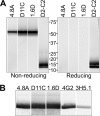
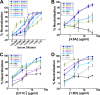
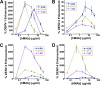
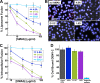
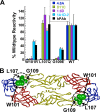
There are no available vaccines for dengue, the most important mosquito-transmitted viral disease. Mechanistic studies with anti-dengue virus (DENV) human monoclonal antibodies (hMAbs) provide a rational approach to identify and characterize neutralizing epitopes on DENV structural proteins that can serve to inform vaccine strategies. Here, we report a class of hMAbs that is likely to be an important determinant in the human humoral response to DENV infection. In this study, we identified and characterized three broadly neutralizing anti-DENV hMAbs: 4.8A, D11C, and 1.6D. These antibodies were isolated from three different convalescent patients with distinct histories of DENV infection yet demonstrated remarkable similarities. All three hMAbs recognized the E glycoprotein with high affinity, neutralized all four serotypes of DENV, and mediated antibody-dependent enhancement of infection in Fc receptor-bearing cells at subneutralizing concentrations. The neutralization activities of these hMAbs correlated with a strong inhibition of virus-liposome and intracellular fusion, not virus-cell binding. We mapped epitopes of these antibodies to the highly conserved fusion loop region of E domain II. Mutations at fusion loop residues W101, L107, and/or G109 significantly reduced the binding of the hMAbs to E protein. The results show that hMAbs directed against the highly conserved E protein fusion loop block viral entry downstream of virus-cell binding by inhibiting E protein-mediated fusion. Characterization of hMAbs targeting this region may provide new insights into DENV vaccine and therapeutic strategies.
Figures

Similar articles
-
The potent and broadly neutralizing human dengue virus-specific monoclonal antibody 1C19 reveals a unique cross-reactive epitope on the bc loop of domain II of the envelope protein.mBio. 2013 Nov 19;4(6):e00873-13. doi: 10.1128/mBio.00873-13. mBio. 2013. PMID: 24255124 Free PMC article.
-
Neutralizing and non-neutralizing monoclonal antibodies against dengue virus E protein derived from a naturally infected patient.Virol J. 2010 Feb 4;7:28. doi: 10.1186/1743-422X-7-28. Virol J. 2010. PMID: 20132551 Free PMC article.
-
Isolation of dengue virus-specific memory B cells with live virus antigen from human subjects following natural infection reveals the presence of diverse novel functional groups of antibody clones.J Virol. 2014 Nov;88(21):12233-41. doi: 10.1128/JVI.00247-14. Epub 2014 Aug 6. J Virol. 2014. PMID: 25100837 Free PMC article.
-
Neutralizing human monoclonal antibodies to severe acute respiratory syndrome coronavirus: target, mechanism of action, and therapeutic potential.Rev Med Virol. 2012 Jan;22(1):2-17. doi: 10.1002/rmv.706. Epub 2011 Sep 8. Rev Med Virol. 2012. PMID: 21905149 Free PMC article. Review.
-
The human antibody response to dengue virus infection.Viruses. 2011 Dec;3(12):2374-95. doi: 10.3390/v3122374. Epub 2011 Nov 25. Viruses. 2011. PMID: 22355444 Free PMC article. Review.
Cited by
-
Evidence of RedOX Imbalance during Zika Virus Infection Promoting the Formation of Disulfide-Bond-Dependent Oligomers of the Envelope Protein.Viruses. 2022 May 24;14(6):1131. doi: 10.3390/v14061131. Viruses. 2022. PMID: 35746600 Free PMC article.
-
The potent and broadly neutralizing human dengue virus-specific monoclonal antibody 1C19 reveals a unique cross-reactive epitope on the bc loop of domain II of the envelope protein.mBio. 2013 Nov 19;4(6):e00873-13. doi: 10.1128/mBio.00873-13. mBio. 2013. PMID: 24255124 Free PMC article.
-
DENV-2 subunit proteins fused to CR2 receptor-binding domain (P28)-induces specific and neutralizing antibodies to the Dengue virus in mice.Hum Vaccin Immunother. 2013 Nov;9(11):2326-35. doi: 10.4161/hv.25673. Epub 2013 Jul 23. Hum Vaccin Immunother. 2013. PMID: 23880886 Free PMC article.
-
Potent Neutralizing Human Monoclonal Antibodies Preferentially Target Mature Dengue Virus Particles: Implication for Novel Strategy for Dengue Vaccine.J Virol. 2018 Nov 12;92(23):e00556-18. doi: 10.1128/JVI.00556-18. Print 2018 Dec 1. J Virol. 2018. PMID: 30185598 Free PMC article.
-
Molecular basis of dengue virus serotype 2 morphological switch from 29°C to 37°C.PLoS Pathog. 2019 Sep 19;15(9):e1007996. doi: 10.1371/journal.ppat.1007996. eCollection 2019 Sep. PLoS Pathog. 2019. PMID: 31536610 Free PMC article.
References
-
- Mackenzie JS, Gubler DJ, Petersen LR. 2004. Emerging flaviviruses: the spread and resurgence of Japanese encephalitis, West Nile and dengue viruses. Nat. Med. 10:S98–S109 - PubMed
-
- WHO 2012. Dengue and severe dengue. Fact sheet no. 117. http://www.who.int/mediacentre/factsheets/fs117/en/
-
- Gubler DJ. 2002. Epidemic dengue/dengue hemorrhagic fever as a public health, social and economic problem in the 21st century. Trends Microbiol. 10:100–103 - PubMed
-
- CDC 2010. Locally acquired dengue—Key West, Florida, 2009–2010. MMWR Morb. Mortal. Wkly. Rep. 59:577–581 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

