Single-molecule photon stamping FRET spectroscopy study of enzymatic conformational dynamics
- PMID: 23085845
- PMCID: PMC3657739
- DOI: 10.1039/c2cp42944f
Single-molecule photon stamping FRET spectroscopy study of enzymatic conformational dynamics
Abstract
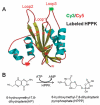
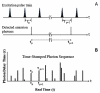
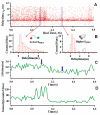
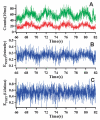
The fluorescence resonant energy transfer (FRET) from a donor to an acceptor via transition dipole-dipole interactions decreases the donor's fluorescent lifetime. The donor's fluorescent lifetime decreases as the FRET efficiency increases, following the equation: E(FRET) = 1 - τ(DA)/τ(D), where τ(D) and τ(DA) are the donor fluorescence lifetime without FRET and with FRET. Accordingly, the FRET time trajectories associated with single-molecule conformational dynamics can be recorded by measuring the donor's lifetime fluctuations. In this article, we report our work on the use of a Cy3/Cy5-labeled enzyme, HPPK to demonstrate probing single-molecule conformational dynamics in an enzymatic reaction by measuring single-molecule FRET donor lifetime time trajectories. Compared with single-molecule fluorescence intensity-based FRET measurements, single-molecule lifetime-based FRET measurements are independent of fluorescence intensity. The latter has an advantage in terms of eliminating the analysis background noise from the acceptor fluorescence detection leak through noise, excitation light intensity noise, or light scattering noise due to local environmental factors, for example, in a AFM-tip correlated single-molecule FRET measurements. Furthermore, lifetime-based FRET also supports simultaneous single-molecule fluorescence anisotropy.
Figures

Similar articles
-
Probing protein multidimensional conformational fluctuations by single-molecule multiparameter photon stamping spectroscopy.J Phys Chem B. 2014 Oct 16;118(41):11943-55. doi: 10.1021/jp5081498. Epub 2014 Oct 3. J Phys Chem B. 2014. PMID: 25222115 Free PMC article.
-
Manipulating protein conformations by single-molecule AFM-FRET nanoscopy.ACS Nano. 2012 Feb 28;6(2):1221-9. doi: 10.1021/nn2038669. Epub 2012 Feb 1. ACS Nano. 2012. PMID: 22276737 Free PMC article.
-
Manipulating and probing enzymatic conformational fluctuations and enzyme-substrate interactions by single-molecule FRET-magnetic tweezers microscopy.Phys Chem Chem Phys. 2014 Jul 14;16(26):13052-8. doi: 10.1039/c4cp01454e. Phys Chem Chem Phys. 2014. PMID: 24853252
-
Fluorescence resonance energy transfer (FRET) and competing processes in donor-acceptor substituted DNA strands: a comparative study of ensemble and single-molecule data.J Biotechnol. 2002 Jan;82(3):211-31. doi: 10.1016/s1389-0352(01)00039-3. J Biotechnol. 2002. PMID: 11999691 Review.
-
Revealing time bunching effect in single-molecule enzyme conformational dynamics.Phys Chem Chem Phys. 2011 Apr 21;13(15):6734-49. doi: 10.1039/c0cp02860f. Epub 2011 Mar 15. Phys Chem Chem Phys. 2011. PMID: 21409227 Review.
Cited by
-
Probing protein multidimensional conformational fluctuations by single-molecule multiparameter photon stamping spectroscopy.J Phys Chem B. 2014 Oct 16;118(41):11943-55. doi: 10.1021/jp5081498. Epub 2014 Oct 3. J Phys Chem B. 2014. PMID: 25222115 Free PMC article.
-
Single-molecule spectroscopy reveals how calmodulin activates NO synthase by controlling its conformational fluctuation dynamics.Proc Natl Acad Sci U S A. 2015 Sep 22;112(38):11835-40. doi: 10.1073/pnas.1508829112. Epub 2015 Aug 26. Proc Natl Acad Sci U S A. 2015. PMID: 26311846 Free PMC article.
-
Fluorescent Organic Small Molecule Probes for Bioimaging and Detection Applications.Molecules. 2022 Dec 1;27(23):8421. doi: 10.3390/molecules27238421. Molecules. 2022. PMID: 36500513 Free PMC article. Review.
-
Conformational states and fluctuations in endothelial nitric oxide synthase under calmodulin regulation.Biophys J. 2021 Dec 7;120(23):5196-5206. doi: 10.1016/j.bpj.2021.11.001. Epub 2021 Nov 6. Biophys J. 2021. PMID: 34748763 Free PMC article.
-
Probing conformational dynamics of an enzymatic active site by an in situ single fluorogenic probe under piconewton force manipulation.Proc Natl Acad Sci U S A. 2016 Dec 27;113(52):15006-15011. doi: 10.1073/pnas.1613404114. Epub 2016 Dec 8. Proc Natl Acad Sci U S A. 2016. PMID: 27940917 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

