Wig1 prevents cellular senescence by regulating p21 mRNA decay through control of RISC recruitment
- PMID: 23085987
- PMCID: PMC3501221
- DOI: 10.1038/emboj.2012.286
Wig1 prevents cellular senescence by regulating p21 mRNA decay through control of RISC recruitment
Abstract
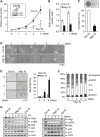
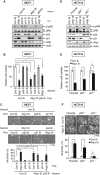
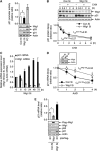
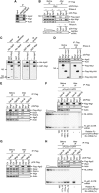
Premature senescence, a key strategy used to suppress carcinogenesis, can be driven by p53/p21 proteins in response to various stresses. Here, we demonstrate that Wig1 plays a critical role in this process through regulation of p21 mRNA stability. Wig1 controls the association of Argonaute2 (Ago2), a central component of the RNA-induced silencing complex (RISC), with target p21 mRNA via binding of the stem-loop structure near the microRNA (miRNA) target site. Depletion of Wig1 prohibited miRNA-mediated p21 mRNA decay and resulted in premature senescence. Wig1 plays an essential role in cell proliferation, as demonstrated in tumour xenografts in mice, and Wig1 and p21 mRNA levels are inversely correlated in human normal and cancer tissues. Together, our data indicate a novel role of Wig1 in RISC target accessibility, which is a key step in RNA-mediated gene silencing. In addition, these findings indicate that fine-tuning of p21 levels by Wig1 is essential for the prevention of cellular senescence.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures


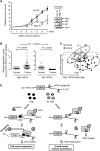
Similar articles
-
WIG1 is crucial for AGO2-mediated ACOT7 mRNA silencing via miRNA-dependent and -independent mechanisms.Nucleic Acids Res. 2017 Jun 20;45(11):6894-6910. doi: 10.1093/nar/gkx307. Nucleic Acids Res. 2017. PMID: 28472401 Free PMC article.
-
Splicing factor SF3B4 acts as a switch in cancer cell senescence by regulating p21 mRNA stability.Cancer Lett. 2025 Apr 10;615:217530. doi: 10.1016/j.canlet.2025.217530. Epub 2025 Feb 15. Cancer Lett. 2025. PMID: 39961431
-
RNA-Binding Protein FXR1 Regulates p21 and TERC RNA to Bypass p53-Mediated Cellular Senescence in OSCC.PLoS Genet. 2016 Sep 8;12(9):e1006306. doi: 10.1371/journal.pgen.1006306. eCollection 2016 Sep. PLoS Genet. 2016. PMID: 27606879 Free PMC article.
-
Rbm24, an RNA-binding protein and a target of p53, regulates p21 expression via mRNA stability.J Biol Chem. 2014 Feb 7;289(6):3164-75. doi: 10.1074/jbc.M113.524413. Epub 2013 Dec 19. J Biol Chem. 2014. PMID: 24356969 Free PMC article.
-
A novel long noncoding RNA Linc-ASEN represses cellular senescence through multileveled reduction of p21 expression.Cell Death Differ. 2020 Jun;27(6):1844-1861. doi: 10.1038/s41418-019-0467-6. Epub 2019 Dec 9. Cell Death Differ. 2020. PMID: 31819156 Free PMC article.
Cited by
-
The p53 mRNA: an integral part of the cellular stress response.Nucleic Acids Res. 2019 Apr 23;47(7):3257-3271. doi: 10.1093/nar/gkz124. Nucleic Acids Res. 2019. PMID: 30828720 Free PMC article. Review.
-
Oncogenic role of rab escort protein 1 through EGFR and STAT3 pathway.Cell Death Dis. 2017 Feb 23;8(2):e2621. doi: 10.1038/cddis.2017.50. Cell Death Dis. 2017. PMID: 28230863 Free PMC article.
-
Heparan sulfation is essential for the prevention of cellular senescence.Cell Death Differ. 2016 Mar;23(3):417-29. doi: 10.1038/cdd.2015.107. Epub 2015 Aug 7. Cell Death Differ. 2016. PMID: 26250908 Free PMC article.
-
Genome-wide identification of Wig-1 mRNA targets by RIP-Seq analysis.Oncotarget. 2016 Jan 12;7(2):1895-911. doi: 10.18632/oncotarget.6557. Oncotarget. 2016. PMID: 26672765 Free PMC article.
-
Roquin binds microRNA-146a and Argonaute2 to regulate microRNA homeostasis.Nat Commun. 2015 Feb 20;6:6253. doi: 10.1038/ncomms7253. Nat Commun. 2015. PMID: 25697406 Free PMC article.
References
-
- Bhattacharyya SN, Habermacher R, Martine U, Closs EI, Filipowicz W (2006) Relief of microRNA-mediated translational repression in human cells subjected to stress. Cell 125: 1111–1124 - PubMed
-
- Brodersen P, Voinnet O (2009) Revisiting the principles of microRNA target recognition and mode of action. Nat Rev Mol Cell Biol 10: 141–148 - PubMed
-
- Brugarolas J, Chandrasekaran C, Gordon JI, Beach D, Jacks T, Hannon GJ (1995) Radiation-induced cell cycle arrest compromised by p21 deficiency. Nature 377: 552–557 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

