New functions for the ancient DedA membrane protein family
- PMID: 23086209
- PMCID: PMC3536176
- DOI: 10.1128/JB.01006-12
New functions for the ancient DedA membrane protein family
Abstract
The DedA protein family is a highly conserved and ancient family of membrane proteins with representatives in most sequenced genomes, including those of bacteria, archaea, and eukarya. The functions of the DedA family proteins remain obscure. However, recent genetic approaches have revealed important roles for certain bacterial DedA family members in membrane homeostasis. Bacterial DedA family mutants display such intriguing phenotypes as cell division defects, temperature sensitivity, altered membrane lipid composition, elevated envelope-related stress responses, and loss of proton motive force. The DedA family is also essential in at least two species of bacteria: Borrelia burgdorferi and Escherichia coli. Here, we describe the phylogenetic distribution of the family and summarize recent progress toward understanding the functions of the DedA membrane protein family.
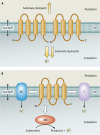
Figures






Similar articles
-
Multiple envelope stress response pathways are activated in an Escherichia coli strain with mutations in two members of the DedA membrane protein family.J Bacteriol. 2013 Jan;195(1):12-24. doi: 10.1128/JB.00762-12. Epub 2012 Oct 5. J Bacteriol. 2013. PMID: 23042993 Free PMC article.
-
Escherichia coli YqjA, a Member of the Conserved DedA/Tvp38 Membrane Protein Family, Is a Putative Osmosensing Transporter Required for Growth at Alkaline pH.J Bacteriol. 2015 Jul;197(14):2292-300. doi: 10.1128/JB.00175-15. Epub 2015 Apr 27. J Bacteriol. 2015. PMID: 25917916 Free PMC article.
-
BB0250 of Borrelia burgdorferi is a conserved and essential inner membrane protein required for cell division.J Bacteriol. 2010 Dec;192(23):6105-15. doi: 10.1128/JB.00571-10. Epub 2010 Sep 24. J Bacteriol. 2010. PMID: 20870761 Free PMC article.
-
Reversing transmembrane electron flow: the DsbD and DsbB protein families.J Mol Microbiol Biotechnol. 2003;5(3):133-49. doi: 10.1159/000070263. J Mol Microbiol Biotechnol. 2003. PMID: 12766342 Review.
-
Metal homeostasis in bacteria: the role of ArsR-SmtB family of transcriptional repressors in combating varying metal concentrations in the environment.Biometals. 2017 Aug;30(4):459-503. doi: 10.1007/s10534-017-0020-3. Epub 2017 May 16. Biometals. 2017. PMID: 28512703 Review.
Cited by
-
Comparative phylogenomics of pathogenic and non-pathogenic mycobacterium.PLoS One. 2013 Aug 28;8(8):e71248. doi: 10.1371/journal.pone.0071248. eCollection 2013. PLoS One. 2013. PMID: 24015186 Free PMC article.
-
Regulation of ER-derived membrane dynamics by the DedA domain-containing proteins VMP1 and TMEM41B.EMBO Rep. 2022 Feb 3;23(2):e53894. doi: 10.15252/embr.202153894. Epub 2022 Jan 19. EMBO Rep. 2022. PMID: 35044051 Free PMC article. Review.
-
A DedA Family Membrane Protein in Indium Extrusion in Rhodanobacter sp. B2A1Ga4.Front Microbiol. 2021 Nov 26;12:772127. doi: 10.3389/fmicb.2021.772127. eCollection 2021. Front Microbiol. 2021. PMID: 34925279 Free PMC article.
-
Bioinformatic analyses of integral membrane transport proteins encoded within the genome of the planctomycetes species, Rhodopirellula baltica.Biochim Biophys Acta. 2014 Jan;1838(1 Pt B):193-215. doi: 10.1016/j.bbamem.2013.08.007. Epub 2013 Aug 19. Biochim Biophys Acta. 2014. PMID: 23969110 Free PMC article.
-
Oxydifficidin, a potent Neisseria gonorrhoeae antibiotic due to DedA assisted uptake and ribosomal protein RplL sensitivity.bioRxiv [Preprint]. 2024 Sep 25:2024.05.27.596031. doi: 10.1101/2024.05.27.596031. bioRxiv. 2024. Update in: Elife. 2025 May 28;13:RP99281. doi: 10.7554/eLife.99281. PMID: 38854004 Free PMC article. Updated. Preprint.
References
-
- Elofsson A, von Heijne G. 2007. Membrane protein structure: prediction vs reality. Annu. Rev. Biochem. 76: 125–140 - PubMed
-
- White SH. 2009. Biophysical dissection of membrane proteins. Nature 459: 344–346 - PubMed
-
- Nonet ML, Marvel CC, Tolan DR. 1987. The hisT-purF region of the Escherichia coli K-12 chromosome. Identification of additional genes of the hisT and purF operons. J. Biol. Chem. 262: 12209–12217 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

