Genomic characterization of Japanese macaque rhadinovirus, a novel herpesvirus isolated from a nonhuman primate with a spontaneous inflammatory demyelinating disease
- PMID: 23097433
- PMCID: PMC3536378
- DOI: 10.1128/JVI.02194-12
Genomic characterization of Japanese macaque rhadinovirus, a novel herpesvirus isolated from a nonhuman primate with a spontaneous inflammatory demyelinating disease
Abstract
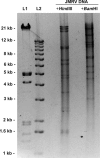
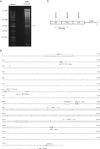
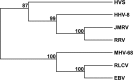
Japanese macaque rhadinovirus (JMRV) is a novel gamma-2 herpesvirus that was isolated from a Japanese macaque (JM) with an inflammatory demyelinating encephalomyelitis referred to as Japanese macaque encephalomyelitis, a disease that possesses clinical and histopathological features resembling multiple sclerosis in humans. Genomic DNA sequence analysis reveals that JMRV is a gammaherpesvirus closely related to rhesus macaque rhadinovirus (RRV) and human herpesvirus 8. We describe here the complete nucleotide sequence and structure of the JMRV genome, as well as the sequence of two plaque isolates of this virus. Analysis of the JMRV genome not only demonstrates that this virus shares a number of genes with RRV that may be involved in pathogenesis but also indicates the presence of unique JMRV genes that could potentially contribute to disease development. The knowledge of the genomic sequence of JMRV, and the ability to easily propagate the virus in vitro, make JMRV infection of JM an attractive model for examining the potential role of an infectious viral agent in the development of demyelinating encephalomyelitis disease in vivo.
Figures


References
-
- Meuth SG, Simon OJ, Grimm A, Melzer N, Herrmann AM, Spitzer P, Landgraf P, Wiendl H. 2008. CNS inflammation and neuronal degeneration is aggravated by impaired CD200-CD200R-mediated macrophage silencing. J. Neuroimmunol. 194:62–69 - PubMed
-
- Petersen TN, Brunak S, von Heijne G, Nielsen H. 2011. SignalP 4.0: discriminating signal peptides from transmembrane regions. Nat. Methods 8:785–786 - PubMed
-
- Bendtsen JD, Jensen LJ, Blom N, Von Heijne G, Brunak S. 2004. Feature-based prediction of non-classical and leaderless protein secretion. PEDS 17:349–356 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources

