DNA-demethylating and anti-tumor activity of synthetic miR-29b mimics in multiple myeloma
- PMID: 23100393
- PMCID: PMC3717964
- DOI: 10.18632/oncotarget.675
DNA-demethylating and anti-tumor activity of synthetic miR-29b mimics in multiple myeloma
Abstract
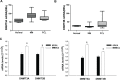
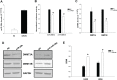
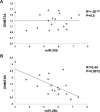
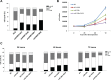
Aberrant DNA methylation plays a relevant role in multiple myeloma (MM) pathogenesis. MicroRNAs (miRNAs) are a class of small non-coding RNAs that recently emerged as master regulator of gene expression by targeting protein-coding mRNAs. However, miRNAs involvement in the regulation of the epigenetic machinery and their potential use as therapeutics in MM remain to be investigated. Here, we provide evidence that the expression of de novo DNA methyltransferases (DNMTs) is deregulated in MM cells. Moreover, we show that miR-29b targets DNMT3A and DNMT3B mRNAs and reduces global DNA methylation in MM cells. In vitro transfection of MM cells with synthetic miR-29b mimics significantly impairs cell cycle progression and also potentiates the growth-inhibitory effects induced by the demethylating agent 5-azacitidine. Most importantly, in vivo intratumor or systemic delivery of synthetic miR-29b mimics, in two clinically relevant murine models of human MM, including the SCID-synth-hu system, induces significant anti-tumor effects. All together, our findings demonstrate that aberrant DNMTs expression is efficiently modulated by tumor suppressive synthetic miR-29b mimics, indicating that methyloma modulation is a novel matter of investigation in miRNA-based therapy of MM.
Conflict of interest statement
The authors declare no competing financial interests.
Figures

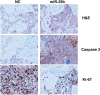
Similar articles
-
Curcumin up-regulates phosphatase and tensin homologue deleted on chromosome 10 through microRNA-mediated control of DNA methylation--a novel mechanism suppressing liver fibrosis.FEBS J. 2014 Jan;281(1):88-103. doi: 10.1111/febs.12574. Epub 2013 Nov 5. FEBS J. 2014. PMID: 24138392
-
Targeted delivery of microRNA-29b by transferrin-conjugated anionic lipopolyplex nanoparticles: a novel therapeutic strategy in acute myeloid leukemia.Clin Cancer Res. 2013 May 1;19(9):2355-67. doi: 10.1158/1078-0432.CCR-12-3191. Epub 2013 Mar 14. Clin Cancer Res. 2013. PMID: 23493348 Free PMC article.
-
Dysregulation of microRNA expression drives aberrant DNA hypermethylation in basal-like breast cancer.Int J Oncol. 2014 Feb;44(2):563-72. doi: 10.3892/ijo.2013.2197. Epub 2013 Nov 29. Int J Oncol. 2014. PMID: 24297604 Free PMC article.
-
Promises and challenges of MicroRNA-based treatment of multiple myeloma.Curr Cancer Drug Targets. 2012 Sep;12(7):838-46. doi: 10.2174/156800912802429355. Curr Cancer Drug Targets. 2012. PMID: 22671926 Free PMC article. Review.
-
MicroRNAs and exosomes: Small molecules with big actions in multiple myeloma pathogenesis.IUBMB Life. 2020 Mar;72(3):314-333. doi: 10.1002/iub.2211. Epub 2019 Dec 11. IUBMB Life. 2020. PMID: 31828868 Review.
Cited by
-
Epigenetic Aberrations in Multiple Myeloma.Cancers (Basel). 2020 Oct 15;12(10):2996. doi: 10.3390/cancers12102996. Cancers (Basel). 2020. PMID: 33076518 Free PMC article. Review.
-
Two Different Serum MiRNA Signatures Correlate with the Clinical Outcome and Histological Subtype in Pleural Malignant Mesothelioma Patients.PLoS One. 2015 Aug 11;10(8):e0135331. doi: 10.1371/journal.pone.0135331. eCollection 2015. PLoS One. 2015. PMID: 26262875 Free PMC article.
-
The role of epigenetics in the biology of multiple myeloma.Blood Cancer J. 2014 May 2;4(5):e207. doi: 10.1038/bcj.2014.29. Blood Cancer J. 2014. PMID: 24786391 Free PMC article. Review.
-
Immune escape of multiple myeloma cells results from low miR29b and the ensuing epigenetic silencing of proteasome genes.Biomark Res. 2024 Apr 23;12(1):43. doi: 10.1186/s40364-024-00592-y. Biomark Res. 2024. PMID: 38654298 Free PMC article.
-
Exosomal miRNAs in the Tumor Microenvironment of Multiple Myeloma.Cells. 2023 Mar 28;12(7):1030. doi: 10.3390/cells12071030. Cells. 2023. PMID: 37048103 Free PMC article. Review.
References
-
- Morgan GJ, Walker BA, Davies FE. The genetic architecture of multiple myeloma. Nat Rev Cancer. 2012;12(5):335–348. - PubMed
-
- Anderson KC, Carrasco RD. Pathogenesis of myeloma. Annual review of pathology. 2011;6:249–274. - PubMed
-
- Fonseca R, Bergsagel PL, Drach J, Shaughnessy J, Gutierrez N, Stewart AK, Morgan G, Van Ness B, Chesi M, Minvielle S, Neri A, Barlogie B, Kuehl WM, Liebisch P, Davies F, Chen-Kiang S, et al. International Myeloma Working Group molecular classification of multiple myeloma: spotlight review. Leukemia. 2009;23(12):2210–2221. - PMC - PubMed
-
- Rossi M, Di Martino MT, Morelli E, Leotta M, Rizzo A, Grimaldi A, Misso G, Tassone P, Caraglia M. Molecular targets for the treatment of multiple myeloma. Curr Cancer Drug Targets. 2012;12(7):757–767. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

